APOBEC3A
| APOBEC3A | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
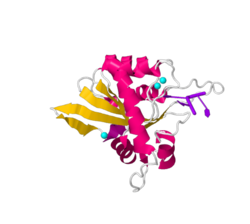 | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | APOBEC3A, A3A, ARP3, PHRBN, bK150C2.1, apolipoprotein B mRNA editing enzyme catalytic subunit 3A | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | OMIM: 607109; HomoloGene: 82288; GeneCards: APOBEC3A; OMA:APOBEC3A - orthologs | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
Apolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 3A, also known as APOBEC3A, or A3A is a gene of the APOBEC3 family found in humans, non-human primates, and some other mammals.[3] It is a single-domain DNA cytidine deaminase with antiviral effects. While other members of the family such as APOBEC3G are believed to act by editing ssDNA by removing an amino group from cytosine in DNA, introducing a cytosine to uracil change which can ultimately lead to a cytosine to thymine mutation, one study suggests that APOBEC3A can inhibit parvoviruses by another mechanism.[4] The cellular function of APOBEC3A is likely to be the destruction of foreign DNA through extensive deamination of cytosine.Stenglein MD, Burns MB, Li M, Lengyel J, Harris RS (February 2010). "APOBEC3 proteins mediate the clearance of foreign DNA from human cells". Nature Structural & Molecular Biology. 17 (2): 222–9. doi:10.1038/nsmb.1744. PMC 2921484. PMID 20062055.
This gene is a member of the polynucleotide cytosine deaminase gene family. It is one of seven related genes or pseudogenes found in a cluster, thought to result from gene duplication, on chromosome 22. Members of the cluster encode proteins that are structurally and functionally related to the C to U RNA-editing cytidine deaminase APOBEC1. The APOBEC3 family of DNA editing enzymes are thought to be part of the innate immune system by restricting retroviruses, mobile genetic elements like retrotransposons and endogenous retroviruses. In addition, APOBEC3A is an important restrictive factor for HIV-1 and other retroviruses such as murine leukemia virus,[5][6]
Structure[edit]
The basic structure APOBEC3A consists of a 5 stranded central β-sheet surrounded by 6 α-helices and a single catalytically active zinc finger domain. Similar to all APOBEC3 catalytic domains, the domain is a HAEx28Cx2-4C zinc binding motif. In such motifs, histidine residues (or cysteine residues in RNA cytidine deaminases) coordinate the zinc ion while a glutamic acid stabilizes the transition state and the proton shuttle. The zinc ion, in this case, is specifically coordinated by residues H70, C101 and C106.[7][8][9][10]
A3A-ssDNA structure[edit]
Single stranded DNA, abbreviated ssDNA, is the substrate that is catalyzed in the C→U deamination reaction of APOBEC3A.
Activity[edit]
A3A has the highest catalytic activity among the APOBEC3 protein family.[11]
mRNA editing activity[edit]
A3A was first found to induce an alternative form of the mRNA editing, G>A, in Wilms' Tumor-1 (WT1) mRNA in cord blood mononuclear cells, particularly in the genomic polymorphic sites, apparently reflecting an amination process rather than a de-amination one.[12] This was soon followed by a study showing A3A induce canonical widespread C>U mRNA editing in human monocytes and macrophages.[13]
Effect of pH on APOBEC3A[edit]
APOBEC3A functions best at an acidic pH, with maximal catalytic activity at pH 5.5.[14][15] Another protein of the APOBEC family very similar to A3A, APOBEC3B, showed little activity at pH 4.5 and 4.0 and a similar assumption can be made of A3A activity at these lower pH levels.[15]
A3A affinity for ssDNA is also pH dependent and closely correlated to the deamination activity of APOBEC3A. The enzyme has the highest affinity for ssDNA at pH 5.5 demonstrating that A3A's maximal catalytic activity and highest affinity for ssDNA occur at a similar pH.[16]
Mechanism of Action[edit]
A3A has become an increasingly widely studied A3 because of its high catalytic activity compared to its family members and its relatively unknown mechanisms compared to more popular APOBEC3's such as APOBEC3G.
Context dependent binding to ssDNA[edit]
The binding of APOBEC3A to its substrate ssDNA is highly dependent on its surrounding nucleotides. The specificity for binding to its target deoxycytidine increases more than ten-fold when the target deoxycytidine is surrounded by deoxythymidine nucleotides.[16]
References[edit]
- ^ a b c ENSG00000262156 GRCh38: Ensembl release 89: ENSG00000128383, ENSG00000262156 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Entrez Gene: APOBEC3A apolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 3A".
- ^ Narvaiza I, Linfesty DC, Greener BN, Hakata Y, Pintel DJ, Logue E, et al. (May 2009). Jung JU (ed.). "Deaminase-independent inhibition of parvoviruses by the APOBEC3A cytidine deaminase". PLOS Pathogens. 5 (5): e1000439. doi:10.1371/journal.ppat.1000439. PMC 2678267. PMID 19461882.
- ^ Jaguva Vasudevan AA, Becker D, Luedde T, Gohlke H, Münk C (March 2021). "Foamy Viruses, Bet, and APOBEC3 Restriction". Viruses. 13 (3): 504. doi:10.3390/v13030504. PMC 8003144. PMID 33803830.
- ^ Stupfler B, Verriez C, Gallois-Montbrun S, Marquet R, Paillart JC (April 2021). "Degradation-Independent Inhibition of APOBEC3G by the HIV-1 Vif Protein". Viruses. 13 (4): 617. doi:10.3390/v13040617. PMC 8066197. PMID 33916704.
- ^ Kouno T, Silvas TV, Hilbert BJ, Shandilya SM, Bohn MF, Kelch BA, et al. (April 2017). "Crystal structure of APOBEC3A bound to single-stranded DNA reveals structural basis for cytidine deamination and specificity". Nature Communications. 8 (1): 15024. Bibcode:2017NatCo...815024K. doi:10.1038/ncomms15024. PMC 5414352. PMID 28452355.
- ^ Byeon IJ, Ahn J, Mitra M, Byeon CH, Hercík K, Hritz J, et al. (2013). "NMR structure of human restriction factor APOBEC3A reveals substrate binding and enzyme specificity". Nature Communications. 4: 1890. Bibcode:2013NatCo...4.1890B. doi:10.1038/ncomms2883. PMC 3674325. PMID 23695684.
- ^ McDaniel YZ, Wang D, Love RP, Adolph MB, Mohammadzadeh N, Chelico L, Mansky LM (February 2020). "Deamination hotspots among APOBEC3 family members are defined by both target site sequence context and ssDNA secondary structure". Nucleic Acids Research. 48 (3): 1353–1371. doi:10.1093/nar/gkz1164. PMC 7026630. PMID 31943071.
- ^ Bohn MF, Shandilya SM, Silvas TV, Nalivaika EA, Kouno T, Kelch BA, et al. (May 2015). "The ssDNA Mutator APOBEC3A Is Regulated by Cooperative Dimerization". Structure. 23 (5): 903–911. doi:10.1016/j.str.2015.03.016. PMC 4874493. PMID 25914058.
- ^ Carpenter MA, Li M, Rathore A, Lackey L, Law EK, Land AM, et al. (October 2012). "Methylcytosine and normal cytosine deamination by the foreign DNA restriction enzyme APOBEC3A". The Journal of Biological Chemistry. 287 (41): 34801–8. doi:10.1074/jbc.m112.385161. PMC 3464582. PMID 22896697.
- ^ Niavarani A, Currie E, Reyal Y, Anjos-Afonso F, Horswell S, Griessinger E, et al. (2015-03-25). "APOBEC3A is implicated in a novel class of G-to-A mRNA editing in WT1 transcripts". PLOS ONE. 10 (3): e0120089. Bibcode:2015PLoSO..1020089N. doi:10.1371/journal.pone.0120089. PMC 4373805. PMID 25807502.
- ^ Sharma S, Patnaik SK, Taggart RT, Kannisto ED, Enriquez SM, Gollnick P, Baysal BE (April 2015). "APOBEC3A cytidine deaminase induces RNA editing in monocytes and macrophages". Nature Communications. 6 (1): 6881. Bibcode:2015NatCo...6.6881S. doi:10.1038/ncomms7881. PMC 4411297. PMID 25898173.
- ^ Pham P, Landolph A, Mendez C, Li N, Goodman MF (October 2013). "A biochemical analysis linking APOBEC3A to disparate HIV-1 restriction and skin cancer". The Journal of Biological Chemistry. 288 (41): 29294–304. doi:10.1074/jbc.M113.504175. PMC 3795231. PMID 23979356.
- ^ a b Ito F, Fu Y, Kao SA, Yang H, Chen XS (June 2017). "Family-Wide Comparative Analysis of Cytidine and Methylcytidine Deamination by Eleven Human APOBEC Proteins". Journal of Molecular Biology. 429 (12): 1787–1799. doi:10.1016/j.jmb.2017.04.021. PMC 5530319. PMID 28479091.
- ^ a b Silvas TV, Hou S, Myint W, Nalivaika E, Somasundaran M, Kelch BA, et al. (May 2018). "Substrate sequence selectivity of APOBEC3A implicates intra-DNA interactions". Scientific Reports. 8 (1): 7511. Bibcode:2018NatSR...8.7511S. doi:10.1038/s41598-018-25881-z. PMC 5951847. PMID 29760455.
Further reading[edit]
- Wedekind JE, Dance GS, Sowden MP, Smith HC (April 2003). "Messenger RNA editing in mammals: new members of the APOBEC family seeking roles in the family business". Trends in Genetics. 19 (4): 207–16. doi:10.1016/S0168-9525(03)00054-4. PMID 12683974.
- Rasmussen HH, van Damme J, Puype M, Gesser B, Celis JE, Vandekerckhove J (December 1992). "Microsequences of 145 proteins recorded in the two-dimensional gel protein database of normal human epidermal keratinocytes". Electrophoresis. 13 (12): 960–9. doi:10.1002/elps.11501301199. PMID 1286667. S2CID 41855774.
- Madsen P, Anant S, Rasmussen HH, Gromov P, Vorum H, Dumanski JP, et al. (August 1999). "Psoriasis upregulated phorbolin-1 shares structural but not functional similarity to the mRNA-editing protein apobec-1". The Journal of Investigative Dermatology. 113 (2): 162–9. doi:10.1046/j.1523-1747.1999.00682.x. PMID 10469298.
- Dunham I, Shimizu N, Roe BA, Chissoe S, Hunt AR, Collins JE, et al. (December 1999). "The DNA sequence of human chromosome 22". Nature. 402 (6761): 489–95. Bibcode:1999Natur.402..489D. doi:10.1038/990031. PMID 10591208.
- Jarmuz A, Chester A, Bayliss J, Gisbourne J, Dunham I, Scott J, Navaratnam N (March 2002). "An anthropoid-specific locus of orphan C to U RNA-editing enzymes on chromosome 22". Genomics. 79 (3): 285–96. doi:10.1006/geno.2002.6718. PMID 11863358.
- Izmailova E, Bertley FM, Huang Q, Makori N, Miller CJ, Young RA, Aldovini A (February 2003). "HIV-1 Tat reprograms immature dendritic cells to express chemoattractants for activated T cells and macrophages". Nature Medicine. 9 (2): 191–7. doi:10.1038/nm822. PMID 12539042. S2CID 26145639.
- Mariani R, Chen D, Schröfelbauer B, Navarro F, König R, Bollman B, et al. (July 2003). "Species-specific exclusion of APOBEC3G from HIV-1 virions by Vif". Cell. 114 (1): 21–31. doi:10.1016/S0092-8674(03)00515-4. PMID 12859895.
- Collins JE, Wright CL, Edwards CA, Davis MP, Grinham JA, Cole CG, et al. (2005). "A genome annotation-driven approach to cloning the human ORFeome". Genome Biology. 5 (10): R84. doi:10.1186/gb-2004-5-10-r84. PMC 545604. PMID 15461802.
- Bogerd HP, Wiegand HL, Doehle BP, Lueders KK, Cullen BR (2006). "APOBEC3A and APOBEC3B are potent inhibitors of LTR-retrotransposon function in human cells". Nucleic Acids Research. 34 (1): 89–95. doi:10.1093/nar/gkj416. PMC 1326241. PMID 16407327.
- Goila-Gaur R, Khan MA, Miyagi E, Kao S, Strebel K (August 2007). "Targeting APOBEC3A to the viral nucleoprotein complex confers antiviral activity". Retrovirology. 4: 61. doi:10.1186/1742-4690-4-61. PMC 2018723. PMID 17727729.
- Biayna J, Garcia-Cao I, Álvarez MM, Salvadores M, Espinosa-Carrasco J, McCullough M, Supek F, Stracker TH (2021-03-31). "Loss of the abasic site sensor HMCES is synthetic lethal with the activity of the APOBEC3A cytosine deaminase in cancer cells". PLOS Biology. 19 (3): e3001176. doi:10.1371/journal.pbio.3001176. PMC 8041192. PMID 33788831.


