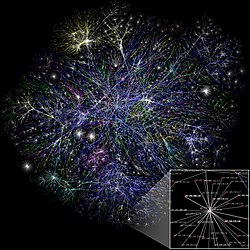
Biological network
| Part of a series on | ||||
| Network science | ||||
|---|---|---|---|---|
| Network types | ||||
| Graphs | ||||
|
||||
| Models | ||||
|
||||
| ||||
This article needs additional citations for verification. (October 2011) |
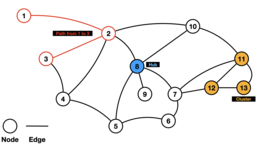
A biological network is a method of representing systems as complex sets of binary interactions or relations between various biological entities.[1] In general, networks or graphs are used to capture relationships between entities or objects.[1] A typical graphing representation consists of a set of nodes connected by edges.
History of networks[edit]

As early as 1736 Leonhard Euler analyzed a real-world issue known as the Seven Bridges of Königsberg, which established the foundation of graph theory. From the 1930's-1950's the study of random graphs were developed. During the mid 1990's, it was discovered that many different types of "real" networks have structural properties quite different from random networks.[2] In the late 2000's, scale-free and small-world networks began shaping the emergence of systems biology, network biology, and network medicine.[1] In 2014, graph theoretical methods were used by Frank Emmert-Streib to analyze biological networks.
In the 1980s, researchers started viewing DNA or genomes as the dynamic storage of a language system with precise computable finite states represented as a finite state machine.[3] Recent complex systems research has also suggested some far-reaching commonality in the organization of information in problems from biology, computer science, and physics.
Networks in biology[edit]
Protein–protein interaction networks[edit]

Protein-protein interaction networks (PINs) represent the physical relationship among proteins present in a cell, where proteins are nodes, and their interactions are undirected edges.[4] Due to their undirected nature, it is difficult to identify all the proteins involved in an interaction. Protein–protein interactions (PPIs) are essential to the cellular processes and also the most intensely analyzed networks in biology. PPIs could be discovered by various experimental techniques, among which the yeast two-hybrid system is a commonly used technique for the study of binary interactions.[5] Recently, high-throughput studies using mass spectrometry have identified large sets of protein interactions.[6]
Many international efforts have resulted in databases that catalog experimentally determined protein-protein interactions. Some of them are the Human Protein Reference Database, Database of Interacting Proteins, the Molecular Interaction Database (MINT),[7] IntAct,[8] and BioGRID.[9] At the same time, multiple computational approaches have been proposed to predict interactions.[10] FunCoup and STRING are examples of such databases, where protein-protein interactions inferred from multiple evidences are gathered and made available for public usage.
Recent studies have indicated the conservation of molecular networks through deep evolutionary time.[11] Moreover, it has been discovered that proteins with high degrees of connectedness are more likely to be essential for survival than proteins with lesser degrees.[12] This observation suggests that the overall composition of the network (not simply interactions between protein pairs) is vital for an organism's overall functioning.
Gene regulatory networks (DNA–protein interaction networks)[edit]

The genome encodes thousands of genes whose products (mRNAs, proteins) are crucial to the various processes of life, such as cell differentiation, cell survival, and metabolism. Genes produce such products through a process called transcription, which is regulated by a class of proteins called transcription factors. For instance, the human genome encodes almost 1,500 DNA-binding transcription factors that regulate the expression of more than 20,000 human genes.[13] The complete set of gene products and the interactions among them constitutes gene regulatory networks (GRN). GRNs regulate the levels of gene products within the cell and in-turn the cellular processes.
GRNs are represented with genes and transcriptional factors as nodes and the relationship between them as edges. These edges are directional, representing the regulatory relationship between the two ends of the edge. For example., the directed edge from gene A to gene B indicates that A regulates the expression of B. Thus, these directional edges can not only represent the promotion of gene regulation but also its inhibition.
GRNs are usually constructed by utilizing the gene regulation knowledge available from databases such as., Reactome and KEGG. High-throughput measurement technologies, such as microarray, RNA-Seq, ChIP-chip, and ChIP-seq, enabled the accumulation of large-scale transcriptomics data, which could help in understanding the complex gene regulation patterns.[14][15]
Gene co-expression networks (transcript–transcript association networks)[edit]
Gene co-expression networks can be perceived as association networks between variables that measure transcript abundances. These networks have been used to provide a system biologic analysis of DNA microarray data, RNA-seq data, miRNA data, etc. weighted gene co-expression network analysis is extensively used to identify co-expression modules and intramodular hub genes.[16] Co-expression modules may correspond to cell types or pathways, while highly connected intramodular hubs can be interpreted as representatives of their respective modules.
Metabolic networks[edit]

Cells break down the food and nutrients into small molecules necessary for cellular processing through a series of biochemical reactions. These biochemical reactions are catalyzed by enzymes. The complete set of all these biochemical reactions in all the pathways represents the metabolic network. Within the metabolic network, the small molecules take the roles of nodes, and they could be either carbohydrates, lipids, or amino acids. The reactions which convert these small molecules from one form to another are represented as edges. It is possible to use network analyses to infer how selection acts on metabolic pathways.[17]
Signaling networks[edit]

Signals are transduced within cells or in between cells and thus form complex signaling networks which plays a key role in the tissue structure. For instance, the MAPK/ERK pathway is transduced from the cell surface to the cell nucleus by a series of protein-protein interactions, phosphorylation reactions, and other events.[18] Signaling networks typically integrate protein–protein interaction networks, gene regulatory networks, and metabolic networks.[19][20] Single cell sequencing technologies allows the extraction of inter-cellular signaling, an example is NicheNet, which allows to modeling intercellular communication by linking ligands to target genes.[21]
Neuronal networks[edit]
The complex interactions in the brain make it a perfect candidate to apply network theory. Neurons in the brain are deeply connected with one another, and this results in complex networks being present in the structural and functional aspects of the brain.[22] For instance, small-world network properties have been demonstrated in connections between cortical regions of the primate brain[23] or during swallowing in humans.[24] This suggests that cortical areas of the brain are not directly interacting with each other, but most areas can be reached from all others through only a few interactions.
Food webs[edit]
All organisms are connected through feeding interactions. If a species eats or is eaten by another species, they are connected in an intricate food web of predator and prey interactions. The stability of these interactions has been a long-standing question in ecology.[25] That is to say if certain individuals are removed, what happens to the network (i.e., does it collapse or adapt)? Network analysis can be used to explore food web stability and determine if certain network properties result in more stable networks. Moreover, network analysis can be used to determine how selective removals of species will influence the food web as a whole.[26] This is especially important considering the potential species loss due to global climate change.
Between-species interaction networks[edit]
In biology, pairwise interactions have historically been the focus of intense study. With the recent advances in network science, it has become possible to scale up pairwise interactions to include individuals of many species involved in many sets of interactions to understand the structure and function of larger ecological networks.[27] The use of network analysis can allow for both the discovery and understanding of how these complex interactions link together within the system's network, a property that has previously been overlooked. This powerful tool allows for the study of various types of interactions (from competitive to cooperative) using the same general framework.[28] For example, plant-pollinator interactions are mutually beneficial and often involve many different species of pollinators as well as many different species of plants. These interactions are critical to plant reproduction and thus the accumulation of resources at the base of the food chain for primary consumers, yet these interaction networks are threatened by anthropogenic change. The use of network analysis can illuminate how pollination networks work and may, in turn, inform conservation efforts.[29] Within pollination networks, nestedness (i.e., specialists interact with a subset of species that generalists interact with), redundancy (i.e., most plants are pollinated by many pollinators), and modularity play a large role in network stability.[29][30] These network properties may actually work to slow the spread of disturbance effects through the system and potentially buffer the pollination network from anthropogenic changes somewhat.[30] More generally, the structure of species interactions within an ecological network can tell us something about the diversity, richness, and robustness of the network.[31] Researchers can even compare current constructions of species interactions networks with historical reconstructions of ancient networks to determine how networks have changed over time.[32] Much research into these complex species interactions networks is highly concerned with understanding what factors (e.g., species richness, connectance, nature of the physical environment) lead to network stability.[33][34]
Within-species interaction networks[edit]
Network analysis provides the ability to quantify associations between individuals, which makes it possible to infer details about the network as a whole at the species and/or population level.[35] One of the most attractive features of the network paradigm would be that it provides a single conceptual framework in which the social organization of animals at all levels (individual, dyad, group, population) and for all types of interaction (aggressive, cooperative, sexual, etc.) can be studied.[36]
Researchers interested in ethology across many taxa, from insects to primates, are starting to incorporate network analysis into their research. Researchers interested in social insects (e.g., ants and bees) have used network analyses better to understand the division of labor, task allocation, and foraging optimization within colonies.[37][38][39] Other researchers are interested in how specific network properties at the group and/or population level can explain individual-level behaviors. Studies have demonstrated how animal social network structure can be influenced by factors ranging from characteristics of the environment to characteristics of the individual, such as developmental experience and personality. At the level of the individual, the patterning of social connections can be an important determinant of fitness, predicting both survival and reproductive success. At the population level, network structure can influence the patterning of ecological and evolutionary processes, such as frequency-dependent selection and disease and information transmission.[40] For instance, a study on wire-tailed manakins (a small passerine bird) found that a male's degree in the network largely predicted the ability of the male to rise in the social hierarchy (i.e., eventually obtain a territory and matings).[41] In bottlenose dolphin groups, an individual's degree and betweenness centrality values may predict whether or not that individual will exhibit certain behaviors, like the use of side flopping and upside-down lobtailing to lead group traveling efforts; individuals with high betweenness values are more connected and can obtain more information, and thus are better suited to lead group travel and therefore tend to exhibit these signaling behaviors more than other group members.[42]
Social network analysis can also be used to describe the social organization within a species more generally, which frequently reveals important proximate mechanisms promoting the use of certain behavioral strategies. These descriptions are frequently linked to ecological properties (e.g., resource distribution). For example, network analyses revealed subtle differences in the group dynamics of two related equid fission-fusion species, Grevy's zebra and onagers, living in variable environments; Grevy's zebras show distinct preferences in their association choices when they fission into smaller groups, whereas onagers do not.[43] Similarly, researchers interested in primates have also utilized network analyses to compare social organizations across the diverse primate order, suggesting that using network measures (such as centrality, assortativity, modularity, and betweenness) may be useful in terms of explaining the types of social behaviors we see within certain groups and not others.[44]
Finally, social network analysis can also reveal important fluctuations in animal behaviors across changing environments. For example, network analyses in female chacma baboons (Papio hamadryas ursinus) revealed important dynamic changes across seasons that were previously unknown; instead of creating stable, long-lasting social bonds with friends, baboons were found to exhibit more variable relationships which were dependent on short-term contingencies related to group-level dynamics as well as environmental variability.[45] Changes in an individual's social network environment can also influence characteristics such as 'personality': for example, social spiders that huddle with bolder neighbors tend to increase also in boldness.[46] This is a very small set of broad examples of how researchers can use network analysis to study animal behavior. Research in this area is currently expanding very rapidly, especially since the broader development of animal-borne tags and computer vision can be used to automate the collection of social associations.[47] Social network analysis is a valuable tool for studying animal behavior across all animal species and has the potential to uncover new information about animal behavior and social ecology that was previously poorly understood.
DNA-DNA chromatin networks[edit]
Within a nucleus, DNA is constantly in motion. Perpetual actions such as genome folding and Cohesin extrusion morph the shape of a genome in real time. The spatial location of strands of chromatin relative to each other plays an important role in the activation or suppression of certain genes. DNA-DNA Chromatin Networks help biologists to understand these interactions by analyzing commonalities amongst different loci. The size of a network can vary significantly, from a few genes to several thousand and thus network analysis can provide vital support in understanding relationships among different areas of the genome. As an example, analysis of spatially similar loci within the organization in a nucleus with Genome Architecture Mapping (GAM) can be used to construct a network of loci with edges representing highly linked genomic regions.
The first graphic showcases the Hist1 region of the mm9 mouse genome with each node representing genomic loci. Two nodes are connected by an edge if their linkage disequilibrium is greater than the average across all 81 genomic windows. The locations of the nodes within the graphic are randomly selected and the methodology of choosing edges yields a, simple to show, but rudimentary graphical representation of the relationships in the dataset. The second visual exemplifies the same information as the previous; However, the network starts with every loci placed sequentially in a ring configuration. It then pulls nodes together using linear interpolation by their linkage as a percentage. The figure illustrates strong connections between the center genomic windows as well as the edge loci at the beginning and end of the Hist1 region.
Modelling biological networks[edit]
Introduction[edit]
To draw useful information from a biological network, an understanding of the statistical and mathematical techniques of identifying relationships within a network is vital. Procedures to identify association, communities, and centrality within nodes in a biological network can provide insight into the relationships of whatever the nodes represent whether they are genes, species, etc. Formulation of these methods transcends disciplines and relies heavily on graph theory, computer science, and bioinformatics.
Association[edit]

There are many different ways to measure the relationships of nodes when analyzing a network. In many cases, the measure used to find nodes that share similarity within a network is specific to the application it is being used. One of the types of measures that biologists utilize is correlation which specifically centers around the linear relationship between two variables.[48] As an example, weighted gene co-expression network analysis uses Pearson correlation to analyze linked gene expression and understand genetics at a systems level.[49] Another measure of correlation is linkage disequilibrium. Linkage disequilibrium describes the non-random association of genetic sequences among loci in a given chromosome.[50] An example of its use is in detecting relationships in GAM data across genomic intervals based upon detection frequencies of certain loci.[51]
Centrality[edit]
The concept of centrality can be extremely useful when analyzing biological network structures. There are many different methods to measure centrality such as betweenness, degree, Eigenvector, and Katz centrality. Every type of centrality technique can provide different insights on nodes in a particular network; However, they all share the commonality that they are to measure the prominence of a node in a network.[52] In 2005, Researchers at Harvard Medical School utilized centrality measures with the yeast protein interaction network. They found that proteins that exhibited high Betweenness centrality were more essential and translated closely to a given protein's evolutionary age.[53]
Communities[edit]

Studying the community structure of a network by subdividing groups of nodes into like-regions can be an integral tool for bioinformatics when exploring data as a network.[54] A food web of The Secaucus High School Marsh exemplifies the benefits of grouping as the relationships between nodes are far easier to analyze with well-made communities. While the first graphic is hard to visualize, the second provides a better view of the pockets of highly connected feeding relationships that would be expected in a food web. The problem of community detection is still an active problem. Scientists and graph theorists continuously discover new ways of sub sectioning networks and thus a plethora of different algorithms exist for creating these relationships.[55] Like many other tools that biologists utilize to understand data with network models, every algorithm can provide its own unique insight and may vary widely on aspects such as accuracy or time complexity of calculation. In 2002, a food web of marine mammals in the Chesapeake Bay was divided into communities by biologists using a community detection algorithm based on neighbors of nodes with high degree centrality. The resulting communities displayed a sizable split in pelagic and benthic organisms.[56] Two very common community detection algorithms for biological networks are the Louvain Method and Leiden Algorithm.
The Louvain method is a greedy algorithm that attempts to maximize modularity, which favors heavy edges within communities and sparse edges between, within a set of nodes. The algorithm starts by each node being in its own community and iteratively being added to the particular node's community that favors a higher modularity.[57][58] Once no modularity increase can occur by joining nodes to a community, a new weighted network is constructed of communities as nodes with edges representing between-community edges and loops representing edges within a community. The process continues until no increase in modularity occurs.[59] While the Louvain Method provides good community detection, there are a few ways that it is limited. By mainly focusing on maximizing a given measure of modularity, it may be led to craft badly connected communities by degrading a model for the sake of maximizing a modularity metric; However, the Louvain Method performs fairly and is can be easy to understand comparatively to many other community detection algorithms.[58]
The Leiden Algorithm expands on the Louvain Method by providing a number of improvements. When joining nodes to a community, only neighborhoods that have been recently changed are considered. This greatly improves the speed of merging nodes. Another optimization is in the refinement phase in-which the algorithm randomly chooses for a node from a set of communities to merge with. This allows for greater depth in choosing communities as Louvain solely focuses on maximizing the modularity that was chosen. The Leiden algorithm, while more complex than Louvain, performs faster with better community detection and can be a valuable tool for identifying groups.[58]
See also[edit]
- List of omics topics in biology
- Biological network inference
- Biostatistics
- Computational biology
- Systems biology
- Weighted correlation network analysis
- Interactome
- Network medicine
- Ecological network
References[edit]
- ^ a b Koutrouli, Mikaela; Karatzas, Evangelos; Paez-Espino, David; Pavlopoulos, Georgios A. (2020). "A Guide to Conquer the Biological Network Era Using Graph Theory". Frontiers in Bioengineering and Biotechnology. 8: 34. doi:10.3389/fbioe.2020.00034. ISSN 2296-4185. PMC 7004966. PMID 32083072.
- ^ Emmert-Streib, Frank; Dehmer, Matthias (2015). "Biological networks: the microscope of the twenty-first century?". Frontiers in Genetics. 6: 307. doi:10.3389/fgene.2015.00307. ISSN 1664-8021. PMC 4602153. PMID 26528327.
- ^ Searls, D. (1993). Artificial intelligence and molecular biology. Cambridge, MA: MIT Press.
- ^ Habibi, Iman; Emamian, Effat S.; Abdi, Ali (2014-01-01). "Quantitative analysis of intracellular communication and signaling errors in signaling networks". BMC Systems Biology. 8: 89. doi:10.1186/s12918-014-0089-z. ISSN 1752-0509. PMC 4255782. PMID 25115405.
- ^ Mashaghi, A.; et al. (2004). "Investigation of a protein complex network". European Physical Journal. 41 (1): 113–121. arXiv:cond-mat/0304207. Bibcode:2004EPJB...41..113M. doi:10.1140/epjb/e2004-00301-0. S2CID 9233932.
- ^ Smits, Arne H.; Vermeulen, Michiel (2016). "Characterizing Protein–Protein Interactions Using Mass Spectrometry: Challenges and Opportunities". Trends in Biotechnology. 34 (10): 825–834. doi:10.1016/j.tibtech.2016.02.014. hdl:2066/161800. ISSN 0167-7799. PMID 26996615.
- ^ Zanzoni, A; Montecchi-Palazzi, L; Quondam, M; Ausiello, G; Helmer-Citterich, M; Cesareni, G (Feb 20, 2002). "MINT: a Molecular INTeraction database". FEBS Letters. 513 (1): 135–40. doi:10.1016/s0014-5793(01)03293-8. PMC 1751541. PMID 11911893.
- ^ Kerrien, S.; Aranda, B.; Breuza, L.; Bridge, A.; Broackes-Carter, F.; Chen, C.; Duesbury, M.; Dumousseau, M.; Feuermann, M.; Hinz, U.; Jandrasits, C.; Jimenez, R. C.; Khadake, J.; Mahadevan, U.; Masson, P.; Pedruzzi, I.; Pfeiffenberger, E.; Porras, P.; Raghunath, A.; Roechert, B.; Orchard, S.; Hermjakob, H. (24 November 2011). "The IntAct molecular interaction database in 2012". Nucleic Acids Research. 40 (D1): D841–D846. doi:10.1093/nar/gkr1088. PMC 3245075. PMID 22121220.
- ^ Oughtred, Rose; Rust, Jennifer; Chang, Christie; Breitkreutz, Bobby-Joe; Stark, Chris; Willems, Andrew; Boucher, Lorrie; Leung, Genie; Kolas, Nadine; Zhang, Frederick; Dolma, Sonam; Coulombe-Huntington, Jasmin; Chatr-aryamontri, Andrew; Dolinski, Kara; Tyers, Mike (2020). "TheBioGRIDdatabase: A comprehensive biomedical resource of curated protein, genetic, and chemical interactions". Protein Science. 30 (1): 187–200. doi:10.1002/pro.3978. ISSN 0961-8368. PMC 7737760. PMID 33070389.
- ^ Jansen, R. (2003). "A Bayesian Networks Approach for Predicting Protein-Protein Interactions from Genomic Data". Science. 302 (5644): 449–453. Bibcode:2003Sci...302..449J. doi:10.1126/science.1087361. ISSN 0036-8075. PMID 14564010. S2CID 5293611.
- ^ Sharan, R.; et al. (2005). "Conserved patterns of protein interaction in multiple species". Proceedings of the National Academy of Sciences of the United States of America. 102 (6): 1974–1979. Bibcode:2005PNAS..102.1974S. doi:10.1073/pnas.0409522102. PMC 548573. PMID 15687504.
- ^ Jeong, H.; et al. (2001). "Lethality and centrality in protein networks". Nature. 411 (6833): 41–42. arXiv:cond-mat/0105306. Bibcode:2001Natur.411...41J. doi:10.1038/35075138. PMID 11333967. S2CID 258942.
- ^ Vaquerizas, J.-M.; et al. (2009). "A census of human transcription factors: function, expression and evolution". Nature Reviews Genetics. 10 (4): 252–263. doi:10.1038/nrg2538. PMID 19274049. S2CID 3207586.
- ^ Jia, Bochao; Xu, Suwa; Xiao, Guanghua; Lamba, Vishal; Liang, Faming (2017). "Learning gene regulatory networks from next generation sequencing data". Biometrics. 73 (4): 1221–1230. doi:10.1111/biom.12682. ISSN 0006-341X. PMC 6258556. PMID 28294287.
- ^ Angelini, Claudia; Costa, Valerio (2014). "Understanding gene regulatory mechanisms by integrating ChIP-seq and RNA-seq data: statistical solutions to biological problems". Frontiers in Cell and Developmental Biology. 2: 51. doi:10.3389/fcell.2014.00051. ISSN 2296-634X. PMC 4207007. PMID 25364758.
- ^ Zheng, Peng-Fei; Chen, Lu-Zhu; Guan, Yao-Zong; Liu, Peng (2021). "Weighted gene co-expression network analysis identifies specific modules and hub genes related to coronary artery disease". Scientific Reports. 11 (1): 6711. Bibcode:2021NatSR..11.6711Z. doi:10.1038/s41598-021-86207-0. ISSN 2045-2322. PMC 7988178. PMID 33758323.
- ^ Proulx, S.R.; et al. (2005). "Network thinking in ecology and evolution". Trends in Ecology and Evolution. 20 (6): 345–353. doi:10.1016/j.tree.2005.04.004. PMID 16701391.
- ^ Cargnello, M.; Roux, P. P. (2011). "Activation and Function of the MAPKs and Their Substrates, the MAPK-Activated Protein Kinases". Microbiology and Molecular Biology Reviews. 75 (1): 50–83. doi:10.1128/MMBR.00031-10. ISSN 1092-2172. PMC 3063353. PMID 21372320.
- ^ Sevimoglu, Tuba; Arga, Kazim Yalcin (2014). "The role of protein interaction networks in systems biomedicine". Computational and Structural Biotechnology Journal. 11 (18): 22–27. doi:10.1016/j.csbj.2014.08.008. ISSN 2001-0370. PMC 4212283. PMID 25379140.
- ^ Arga, K Yalçın; Önsan, Z İlsen; Kırdar, Betül; Ülgen, Kutlu Ö; Nielsen, Jens (2007). "Understanding signaling in yeast: Insights from network analysis". Biotechnology and Bioengineering. 97 (5): 1246–1258. doi:10.1002/bit.21317. ISSN 0006-3592. PMID 17252576. S2CID 38896124.
- ^ Browaeys, Robin; Saelens, Wouter; Saeys, Yvan (February 2020). "NicheNet: modeling intercellular communication by linking ligands to target genes". Nature Methods. 17 (2): 159–162. doi:10.1038/s41592-019-0667-5. PMID 31819264. S2CID 256836571.
- ^ Bullmore, E. & O. Sporns (2009). "Complex brain networks: graph theoretical analysis of structural and functional systems". Nature Reviews Neuroscience. 10 (3): 186–198. doi:10.1038/nrn2575. PMID 19190637. S2CID 205504722.
- ^ Stephan, K.E.; et al. (2000). "Computational analysis of functional connectivity between areas of primate cerebral cortex". Philosophical Transactions of the Royal Society B. 355 (1393): 111–126. doi:10.1098/rstb.2000.0552. PMC 1692715. PMID 10703047.
- ^ Jestrović, Iva; Coyle, James L; Perera, Subashan; Sejdić, Ervin (2016-12-01). "Functional connectivity patterns of normal human swallowing: difference among various viscosity swallows in normal and chin-tuck head positions". Brain Research. 1652: 158–169. doi:10.1016/j.brainres.2016.09.041. ISSN 0006-8993. PMC 5102805. PMID 27693396.
- ^ MacArthur, R.H. (1955). "Fluctuations in animal populations and a measure of community stability". Ecology. 36 (3): 533–536. Bibcode:1955Ecol...36..533M. doi:10.2307/1929601. JSTOR 1929601.
- ^ Dunne, J.A.; et al. (2002). "Network structure and biodiversity loss in food webs: robustness increases with connectance". Ecology Letters. 5 (4): 558–567. Bibcode:2002EcolL...5..558D. doi:10.1046/j.1461-0248.2002.00354.x. S2CID 2114852.
- ^ Bascompte, J. (2009). "Disentangling the web of life". Science. 325 (5939): 416–419. Bibcode:2009Sci...325..416B. doi:10.1126/science.1170749. PMID 19628856. S2CID 2249052.
- ^ Krause, J.; et al. (2009). "Animal social networks: an introduction". Behav. Ecol. Sociobiol. 63 (7): 967–973. doi:10.1007/s00265-009-0747-0. S2CID 24523607.
- ^ a b Memmott, J.; et al. (2004). "Tolerance of pollination networks to species extinctions". Philosophical Transactions of the Royal Society B. 271 (1557): 2605–261. doi:10.1098/rspb.2004.2909. PMC 1691904. PMID 15615687.
- ^ a b Olesen, J.; et al. (2007). "The modularity of pollination networks". PNAS. 104 (50): 19891–19896. Bibcode:2007PNAS..10419891O. doi:10.1073/pnas.0706375104. PMC 2148393. PMID 18056808.
- ^ Campbell, V.; et al. (2011). "Experimental design and the outcome and interpretation of diversity-stability relations". Oikos. 120 (3): 399–408. Bibcode:2011Oikos.120..399C. doi:10.1111/j.1600-0706.2010.18768.x.
- ^ Lotze, H.; et al. (2011). "Historical changes in marine resources, food-web structure and ecosystem functioning in the Adriatic Sea, Mediterranean". Ecosystems. 14 (2): 198–222. Bibcode:2011Ecosy..14..198L. doi:10.1007/s10021-010-9404-8. S2CID 45894582.
- ^ Romanuk, T.; et al. (2010). "Maintenance of positive diversity-stability relations along a gradient of environmental stress". PLOS ONE. 5 (4): e10378. Bibcode:2010PLoSO...510378R. doi:10.1371/journal.pone.0010378. PMC 2860506. PMID 20436913.
- ^ Briand, F. (1983). "Environmental control of food web structure". Ecology. 64 (2): 253–263. Bibcode:1983Ecol...64..253B. doi:10.2307/1937073. JSTOR 1937073.
- ^ Croft, D.P.; et al. (2004). "Social networks in the guppy (Poecilia reticulate)". Philosophical Transactions of the Royal Society B. 271 (Suppl): S516–S519. doi:10.1098/rsbl.2004.0206. PMC 1810091. PMID 15801620.
- ^ Krause, Jens; Lusseau, David; James, Richard (1 May 2009). "Animal social networks: an introduction". Behavioral Ecology and Sociobiology. 63 (7): 967–973. doi:10.1007/s00265-009-0747-0. S2CID 24523607.
- ^ Dornhaus, A.; et al. (2006). "Benefits of recruitment in honey bees: Effects of ecology and colony size in an individual-based model". Behavioral Ecology. 17 (3): 336–344. doi:10.1093/beheco/arj036.
- ^ Linksvayer, T.; et al. (2012). "Developmental evolution in social insects: Regulatory networks from genes to societies". Journal of Experimental Zoology Part B: Molecular and Developmental Evolution. 318 (3): 159–169. Bibcode:2012JEZB..318..159L. doi:10.1002/jez.b.22001. PMID 22544713.
- ^ Mullen, R.; et al. (2009). "A review of ant algorithms". Expert Systems with Applications. 36 (6): 9608–9617. doi:10.1016/j.eswa.2009.01.020.
- ^ Croft, Darren P.; Darden, Safi K.; Wey, Tina W. (2016). "Current directions in animal social networks". Current Opinion in Behavioral Sciences. 12: 52–58. doi:10.1016/j.cobeha.2016.09.001. hdl:10871/23348. S2CID 53195734.
- ^ Ryder, T.B.; et al. (2008). "Social networks in the lek-mating wire-tailed manakin (Pipra filicauda)". Philosophical Transactions of the Royal Society B. 275 (1641): 1367–1374. doi:10.1098/rspb.2008.0205. PMC 2602714. PMID 18381257.
- ^ Lusseau, D. (2007). "Evidence for social role in a dolphin social network". Evolutionary Ecology. 21 (3): 357–366. arXiv:q-bio/0607048. Bibcode:2007EvEco..21..357L. doi:10.1007/s10682-006-9105-0. S2CID 9748737.
- ^ Sundaresan, S.; et al. (2007). "Network metrics reveal differences in social organization between two fission-fusion species, Grevy's zebra and onager". Oecologia. 151 (1): 140–149. Bibcode:2007Oecol.151..140S. doi:10.1007/s00442-006-0553-6. PMID 16964497. S2CID 8104281.
- ^ Kasper, C.; Voelkl, B. (2009). "A social network analysis of primate groups". Primates. 50 (4): 343–356. doi:10.1007/s10329-009-0153-2. PMID 19533270. S2CID 9852394.
- ^ Henzi, S.; et al. (2009). "Cyclicity in the structure of female baboon social networks". Behavioral Ecology and Sociobiology. 63 (7): 1015–1021. doi:10.1007/s00265-009-0720-y. S2CID 6021233.
- ^ Hunt, E.R.; et al. (2018). "Social interactions shape individual and collective personality in social spiders". Proceedings of the Royal Society B. 285 (1886): 20181366. doi:10.1098/rspb.2018.1366. PMC 6158534. PMID 30185649.
- ^ Krause J, Krause S, Arlinghaus R, Psorakis I, Roberts S, Rutz C (2013). "Reality mining of animal social systems". Trends in Ecology and Evolution. 28 (9): 541–551. doi:10.1016/j.tree.2013.06.002. PMID 23856617. Archived from the original on 2023-02-04. Retrieved 2019-12-08.
- ^ Haug, Mark Gerard. "measure of association". Encyclopedia Britannica, Invalid Date, https://www.britannica.com/topic/measure-of-association Archived 2023-02-04 at the Wayback Machine. Accessed 18 April 2022.
- ^ Zhang, Bin and Steve Horvath. A General Framework for Weighted Gene Co-Expression Network Analysis. Statistical Applications in Genetics and Molecular Biology, vol. 4, no. 1, 2005, article 17. https://dibernardo.tigem.it/files/papers/2008/zhangbin-statappsgeneticsmolbio.pdf
- ^ “Linkage Disequilibrium.” Linkage Disequilibrium - ISOGG Wiki, International Society of Genetic Genealogy, https://isogg.org/wiki/Linkage_disequilibrium Archived 2021-06-08 at the Wayback Machine.
- ^ Beagrie, Robert A et al. “Complex multi-enhancer contacts captured by genome architecture mapping.” Nature vol. 543,7646 (2017): 519-524. doi:10.1038/nature21411
- ^ “Centrality Measure.” Centrality Measure - an Overview | ScienceDirect Topics, ScienceDirect, https://www.sciencedirect.com/topics/computer-science/centrality-measure Archived 2022-04-18 at the Wayback Machine.
- ^ Joy MP, Brock A, Ingber DE, Huang S. High-betweenness proteins in the yeast protein interaction network. J Biomed Biotechnol. 2005 Jun 30;2005(2):96-103. doi: 10.1155/JBB.2005.96. PMID 16046814; PMCID: PMC1184047.
- ^ Porter, Mason A et al. “Communities in Networks” Notices of the AMS vol. 56, no. 9(2009): 1082-1097. https://www.ams.org/notices/200909/rtx090901082p.pdf Archived 2021-06-13 at the Wayback Machine
- ^ “Community Detection.” Community Detection - an Overview | ScienceDirect Topics, ScienceDirect, https://www.sciencedirect.com/topics/computer-science/community-detection.
- ^ Girvan, M, and M E J Newman. “Community structure in social and biological networks.” Proceedings of the National Academy of Sciences of the United States of America vol. 99,12 (2002): 7821-6. doi:10.1073/pnas.122653799
- ^ Markovitch, Omer, and Natalio Krasnogor. “Predicting species emergence in simulated complex pre-biotic networks.” PLOS ONE vol. 13,2 e0192871. 15 Feb. 2018, doi:10.1371/journal.pone.0192871
- ^ a b c Traag, V.A., Waltman, L. & van Eck, N.J. From Louvain to Leiden: guaranteeing well-connected communities. Sci Rep 9, 5233 (2019). https://doi.org/10.1038/s41598-019-41695-z Archived 2023-02-04 at the Wayback Machine
- ^ Ozaki, Naoto. Tezuka Hiroshi. Inaba, Mary. “A Simple Acceleration Method for the Louvain Algorithm” International Journal of Computer and Electrical Engineering, vol. 8, no. 3, 3 June 2016, page numbers pp 207-218. http://www.ijcee.org/vol8/927-A023.pdf Archived 2023-02-04 at the Wayback Machine
Books[edit]
- E. Estrada, "The Structure of Complex Networks: Theory and Applications", Oxford University Press, 2011, ISBN 978-0-199-59175-6
- J. Krause, R. James, D. Franks, D. Croft, "Animal Social Networks", Oxford University Press, 2015, ISBN 978-0199679041
External links[edit]
- Networkbio.org, The site of the series of Integrative Network Biology (INB) meetings. For the 2012 event also see www.networkbio.org
- Network Tools and Applications in Biology (NETTAB) workshops.
- Networkbiology.org, NetworkBiology wiki site.
- Linding Lab, Technical University of Denmark (DTU) studies Network Biology and Cellular Information Processing, and is also organizing the Denmark branch of the annual "Integrative Network Biology and Cancer" symposium series.
- NRNB.org, The National Resource for Network Biology. A US National Institute of Health (NIH) Biomedical Technology Research Center dedicated to the study of biological networks.
- Network Repository The first interactive data and network data repository with real-time visual analytics.
- Animal Social Network Repository (ASNR) The first multi-taxonomic repository that collates 790 social networks from more than 45 species, including those of mammals, reptiles, fish, birds, and insects