Haplogroup L0
| Haplogroup L0 | |
|---|---|
| Possible time of origin | 130 to 200 ka[1][2] |
| Possible place of origin | Southern Africa or Southern East Africa |
| Ancestor | L (Mitochondrial Eve) |
| Descendants | L0a'b'f'k, L0d |
| Defining mutations | 263!, 1048, 3516A, 5442, 6185, 9042, 9347, 10589, 12007, 12720[3] |
Haplogroup L0 is a human mitochondrial DNA (mtDNA) haplogroup.
Origin[edit]

L0 is one of two branches from the most recent common ancestor (MRCA) for the shared human maternal lineage. The haplogroup consists of five main branches (L0a, L0b, L0d, L0f, L0k). Four of them were originally classified into L1 subclades, L1a, L1d, L1f and L1k.
In 2014, ancient DNA analysis of a 2,330 year old male forager's skeleton in Southern Africa found that the specimen belonged to the L0d2c1c mtDNA subclade. This maternal haplogroup is today most closely associated with the Ju, a subgroup of the indigenous San people, which points to population continuity in the region.[4] In 2016, a Late Iron Age desiccated mummy from the Tuli region in northern Botswana was also found to belong to haplogroup L0.[5]
Distribution[edit]

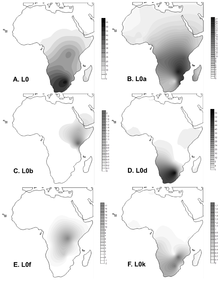
L0 is found most commonly in Sub-Saharan Africa. It reaches its highest frequency in the Khoisan people at 73% on average.[6] Some of the highest frequencies are:[7] Namibia (!Xun) 79%, South Africa (Khwe/!Xun) 83%, and Botswana (!Kung) 100%.
Haplogroup L0d is found among Khoisan groups of Southern Africa closer to the Khoid side with (following L0k) being more Sanid but is largely restricted to the Khoisan as a whole.[7][8][9][10] L0d is also commonly found in sections of the Coloured population of South Africa and frequencies range from 60%[11] to 71%.[10] This illustrates the massive maternal contribution of Khoisan people to sections of the Coloured population of South Africa.
Haplogroups L0k is the second most common haplogroup in the Khoisan groups closer to the Sanid side with (following L0d) being more Khoid but is largely restricted to the Khoisan as a whole.[7][8][9][10] Although the Khoisan associated L0d haplogroup were found in high frequencies in sections of the Coloured population of South Africa, L0k were not observed in two studies involving large groups of Coloured individuals.[10][11]
Haplogroup L0f is present in relatively small frequencies in Tanzania, East Africa among the Sandawe people of Tanzania.
Haplogroup L0a is most prevalent in South-East African populations (25% in Mozambique).[6] Among Guineans, it has a frequency between 1% and 5%, with the Balanta group showing increased frequency of about 11%. Haplogroup L0a has a Paleolithic time depth of about 33,000 years and likely reached Guinea between 10,000 and 4,000 years ago. It also is often seen in the Mbuti and Biaka Pygmies. L0a is found at a frequency of almost 25% in Hadramawt (Yemen).[12]
Haplogroup L0b is found in Ethiopia.
Drug and disease interactions[edit]
In patients who are given the drug stavudine to treat HIV, Haplogroup L0a2 is associated with a higher likelihood of peripheral neuropathy as a side effect.[13]
Subclades[edit]
Tree[edit]

This phylogenetic tree of haplogroup L0 subclades is based on the paper by Mannis van Oven and Manfred Kayser Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation[3] and subsequent published research.
- Most Recent Common Ancestor (MRCA)
- L0
- L0d
- L0d3
- L0d1'2
- L0d1
- L0d1a
- L0d1b
- L0d1c
- L0d1c1
- L0d2
- L0d2a'b
- L0d2a
- L0d2a1
- L0d2b
- L0d2a
- L0d2c
- L0d2c1c[14]
- L0d2a'b
- L0d1
- L0a'b'f'k
- L0k
- L0k1
- L0k2
- L0a'b'f
- L0f
- L0f1
- L0f2
- L0f2a
- L0f2b
- L0a'b
- L0a
- L0a1
- L0a1a
- L0a1a2
- L0a1b
- L0a1b1
- L0a1b1a
- L0a1b2
- L0a1b1
- L0a1c
- L0a1d
- L0a1a
- L0a2
- L0a2a
- L0a2a1
- L0a2a1a
- L0a2a1a1
- L0a2a1a2
- L0a2a1a
- L0a2a2
- L0a2a2a
- L0a2a1
- L0a2b
- L0a2ba
- L0a2c
- L0a2d
- L0a2a
- L0a3
- L0a4
- L0a1
- L0b
- L0a
- L0f
- L0k
- L0d
- L0
See also[edit]
- Genealogical DNA test
- Genetic genealogy
- Human mitochondrial genetics
- Population genetics
- Human mitochondrial DNA haplogroups
|
Phylogenetic tree of human mitochondrial DNA (mtDNA) haplogroups | |||||||||||||||||||||||||||||||||||||||
| Mitochondrial Eve (L) | |||||||||||||||||||||||||||||||||||||||
| L0 | L1–6 | ||||||||||||||||||||||||||||||||||||||
| L1 | L2 | L3 | L4 | L5 | L6 | ||||||||||||||||||||||||||||||||||
| M | N | ||||||||||||||||||||||||||||||||||||||
| CZ | D | E | G | Q | O | A | S | R | I | W | X | Y | |||||||||||||||||||||||||||
| C | Z | B | F | R0 | pre-JT | P | U | ||||||||||||||||||||||||||||||||
| HV | JT | K | |||||||||||||||||||||||||||||||||||||
| H | V | J | T | ||||||||||||||||||||||||||||||||||||
References[edit]
- ^ point estimate 168.5 ka (136.3–201.1 ka 95% CI) according to Heinz, Tanja; et al. (2017). "Updating the African human mitochondrial DNA tree: Relevance to forensic and population genetics". Forensic Science International: Genetics. 27: 156–159. doi:10.1016/j.fsigen.2016.12.016. PMID 28086175. (table 2). 150 ka suggested in:Soares, Pedro; Ermini, Luca; Thomson, Noel; Mormina, Maru; Rito, Teresa; Röhl, Arne; Salas, Antonio; Oppenheimer, Stephen; MacAulay, Vincent (2009). "Correcting for Purifying Selection: An Improved Human Mitochondrial Molecular Clock". The American Journal of Human Genetics. 84 (6): 740–59. doi:10.1016/j.ajhg.2009.05.001. PMC 2694979. PMID 19500773..
- ^ Age estimates (ka, 95% CI in angular brackets): ML whole-mtDNA age estimate: 128.2 [95% CI: 107.9-148.9], ρ whole-mtDNA age estimate: 121.3 [99.2;143.7], ρ synonymous age estimate (ka): 131.0 [97.8;164.2]: Rito T, Richards MB, Fernandes V, Alshamali F, Cerny V, Pereira L, Soares P., "The first modern human dispersals across Africa", PLoS One 2013 Nov 13; 8(11):e80031. doi: 10.1371/journal.pone.0080031.
- ^ a b Van Oven, Mannis; Kayser, Manfred (2009). "Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation". Human Mutation. 30 (2): E386–94. doi:10.1002/humu.20921. PMID 18853457. S2CID 27566749.
- ^ Alan G. Morris; Anja Heinze; Eva K.F. Chan; Andrew B. Smith; Vanessa M. Hayes (2014). "First Ancient Mitochondrial Human Genome from a Pre-Pastoralist Southern African". Genome Biology and Evolution. 6 (10): 2647–53. doi:10.1093/gbe/evu202. PMC 4224329. PMID 25212860.
- ^ Frank J. Rühli; Maryna Steyn; Morongwa N. Mosothwane; Lena Öhrström; Molebogeng K. Bodiba; Abigail Bouwman (January–February 2016). "Radiological and genetic analysis of a Late Iron Age mummy from the Tuli Block, Botswana" (PDF). South African Journal of Science. 112 (1/2). Archived from the original (PDF) on 21 June 2016. Retrieved 26 April 2016.
- ^ a b Rosa, Alexandra; Brehm, Antonio; Kivisild, Toomas; Metspalu, Ene; Villems, Richard (2004). "MtDNA Profile of West Africa Guineans: Towards a Better Understanding of the Senegambia Region". Annals of Human Genetics. 68 (4): 340–52. doi:10.1046/j.1529-8817.2004.00100.x. hdl:10400.13/3044. PMID 15225159. S2CID 15391342.
- ^ a b c Tishkoff, S. A.; Gonder, M. K.; Henn, B. M.; Mortensen, H.; Knight, A.; Gignoux, C.; Fernandopulle, N.; Lema, G.; Nyambo, T. B. (2007). "History of Click-Speaking Populations of Africa Inferred from mtDNA and Y Chromosome Genetic Variation". Molecular Biology and Evolution. 24 (10): 2180–95. doi:10.1093/molbev/msm155. PMID 17656633.
- ^ a b Chen, Yu-Sheng; Olckers, Antonel; Schurr, Theodore G.; Kogelnik, Andreas M.; Huoponen, Kirsi; Wallace, Douglas C. (2000). "MtDNA Variation in the South African Kung and Khwe—and Their Genetic Relationships to Other African Populations". The American Journal of Human Genetics. 66 (4): 1362–83. doi:10.1086/302848. PMC 1288201. PMID 10739760.
- ^ a b Knight, Alec; Underhill, Peter A.; Mortensen, Holly M.; Zhivotovsky, Lev A.; Lin, Alice A.; Henn, Brenna M.; Louis, Dorothy; Ruhlen, Merritt; Mountain, Joanna L. (2003). "African Y Chromosome and mtDNA Divergence Provides Insight into the History of Click Languages". Current Biology. 13 (6): 464–73. doi:10.1016/S0960-9822(03)00130-1. PMID 12646128. S2CID 52862939.
- ^ a b c d Schlebusch, Carina M.; Naidoo, Thijessen; Soodyall, Himla (2009). "SNaPshot minisequencing to resolve mitochondrial macro-haplogroups found in Africa". Electrophoresis. 30 (21): 3657–64. doi:10.1002/elps.200900197. PMID 19810027. S2CID 19515426.
- ^ a b Quintana-Murci, Lluis; Harmant, Christine; Quach, Hélène; Balanovsky, Oleg; Zaporozhchenko, Valery; Bormans, Connie; Van Helden, Paul D.; Hoal, Eileen G.; Behar, Doron M. (2010). "Strong Maternal Khoisan Contribution to the South African Coloured Population: A Case of Gender-Biased Admixture". The American Journal of Human Genetics. 86 (4): 611–20. doi:10.1016/j.ajhg.2010.02.014. PMC 2850426. PMID 20346436.
- ^ Rídl, Jakub; Edens, Christopher M.; Černý, Viktor (2009). "Mitochondrial DNA Structure of Yemeni Population: Regional Differences and the Implications for Different Migratory Contributions". The Evolution of Human Populations in Arabia. Vertebrate Paleobiology and Paleoanthropology. pp. 69–78. doi:10.1007/978-90-481-2719-1_5. ISBN 978-90-481-2718-4.
- ^ Kampira E, Kumwenda J, van Oosterhout JJ, Dandara C. Mitochondrial DNA subhaplogroups L0a2 and L2a modify susceptibility to peripheral neuropathy in malawian adults on stavudine containing highly active antiretroviral therapy., J Acquir Immune Defic Syndr. 2013 Aug 15; 63(5):647-52. doi: 10.1097/QAI.0b013e3182968ea5
- ^ First Ancient Mitochondrial Human Genome from a Pre-Pastoralist Southern African
External links[edit]
- General
- Ian Logan's Mitochondrial DNA Site
- Mannis van Oven's Phylotree
- Haplogroup L0
- L0 YFull MTree 1.02.00 (under construction)
- Rosa, Alexandra; Brehm, Antonio; Kivisild, Toomas; Metspalu, Ene; Villems, Richard (2004). "MtDNA Profile of West Africa Guineans: Towards a Better Understanding of the Senegambia Region". Annals of Human Genetics. 68 (4): 340–52. doi:10.1046/j.1529-8817.2004.00100.x. hdl:10400.13/3044. PMID 15225159. S2CID 15391342.
Based on the previous knowledge of African complete sequences paraphyletic clade L1 is split into two monophyletic units L0, capturing previously defined L1a and L1d lineages, and L1 clade that includes L1b and L1c clades…
- Pavesi, A. (2005). "Utility of JC polyomavirus in tracing the pattern of human migrations dating to prehistoric times". Journal of General Virology. 86 (5): 1315–26. doi:10.1099/vir.0.80650-0. PMID 15831942.
The first axis of mtDNA, on the other hand, places the ancestral haplogroup L0 at the extreme left, since it gave rise to one sole lineage…
- Mishmar, Dan; Ruiz-Pesini, Eduardo; Brandon, Martin; Wallace, Douglas C. (2004). "Mitochondrial DNA-like sequences in the nucleus (NUMTs): Insights into our African origins and the mechanism of foreign DNA integration". Human Mutation. 23 (2): 125–33. doi:10.1002/humu.10304. PMID 14722916. S2CID 25109836.
haplogroup L0 mtDNAs, the haplogroup that we had previously concluded lies at the base of the human mtDNA. tree based on phylogenetic analysis…
- Knight, Alec; Underhill, Peter A.; Mortensen, Holly M.; Zhivotovsky, Lev A.; Lin, Alice A.; Henn, Brenna M.; Louis, Dorothy; Ruhlen, Merritt; Mountain, Joanna L. (2003). "African Y Chromosome and mtDNA Divergence Provides Insight into the History of Click Languages". Current Biology. 13 (6): 464–73. doi:10.1016/S0960-9822(03)00130-1. PMID 12646128. S2CID 52862939.
