Monkeypox virus
| Monkeypox virus | |
|---|---|
| |
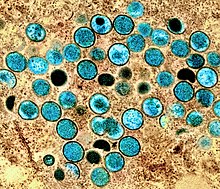
| Colorized transmission electron micrograph of monkeypox virus particles (teal) found within an infected cell (brown), cultured in the laboratory. | |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Varidnaviria |
| Kingdom: | Bamfordvirae |
| Phylum: | Nucleocytoviricota |
| Class: | Pokkesviricetes |
| Order: | Chitovirales |
| Family: | Poxviridae |
| Genus: | Orthopoxvirus |
| Species: | Monkeypox virus
|
| Clades | |
| |
| Synonyms | |
|
MPV, MPXV, hMPXV, [ongoing consideration about changing the virus's name] | |
The monkeypox virus (MPV, MPXV, or hMPXV),[1][a] is a species of double-stranded DNA virus that causes mpox disease in humans and other mammals. The monkeypox virus is a zoonotic virus belonging to the orthopoxvirus genus, making it closely related to the variola, cowpox, and vaccinia viruses. MPV is oval-shaped with a lipoprotein outer membrane. The genome is approximately 190 kb.
The smallpox and monkeypox viruses are both orthopoxviruses, and the smallpox vaccine is effective against mpox if given within 3–5 years before contracting the disease.[3] Symptoms of mpox in humans include a rash that forms blisters and then crusts over, fever, and swollen lymph nodes.[4] The virus is transmissible between animals and humans by direct contact to the lesions or bodily fluids.[5] The virus was given the name monkeypox virus after being isolated from monkeys, but most of the carriers of this virus are small mammals.[4]
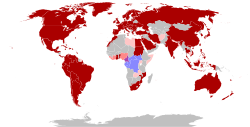
The virus is endemic in Central Africa, where infections in humans are relatively frequent.[4][6] Though there are many natural hosts for the monkeypox virus, the exact reservoirs and how the virus is circulated in nature needs to be studied further.[7]
Virology
Classification
MPV is part of the genus Orthopoxvirus, belonging to the Poxviridae family, which have been listed by the WHO as diseases with epidemic or pandemic potential.[8] There are two subtypes, Clade I historically associated with the Congo Basin and Clade II historically associated with West Africa. A global outbreak during 2022–2023 was caused by Clade II.[4]
MPV is 96.3% identical to the variola virus in regards to its coding region, but it does differ in parts of the genome which encode for virulence and host range.[9] Through phylogenetic analysis, it was found that MPV is not a direct descendant of the variola virus.[9]
Structure and genome
The monkeypox virus, like other poxviruses, is oval shaped, with a lipoprotein outer membrane. The outer membrane protects the enzymes, DNA, and transcription factors of the virus.[10] Typical DNA viruses replicate and express their genome in the nucleus of eukaryotic cells, relying heavily on the host cell's machinery. However, the monkeypox viruses rely mostly on the protein encoded in their genome that allows them to replicate in the cytoplasm.[11]
The genome of the monkeypox virus comprises 200 kb of double stranded DNA coding for 191 proteins.[12][13] Similar to other poxviruses, the virions of monkey pox have large oval shaped envelopes. Within each virion there is a core which holds the genome along with the enzymes that assist in dissolving the protein coat and replication.[14] The center of the genome codes for genes involved in key functions such as viral transcription and assembly; genes located on the extremities of the viral genome are associated more towards interactions between the virus and the host cell such as spike protein characteristics.[11]
Monkeypox virus is relatively large compared to other viruses. This makes it harder for the virus to breach the host defenses, such as crossing past gap junctions. Furthermore, the large size makes it harder for the virus to quickly replicate and evade immune response.[11] To evade host immune systems, and buy more time for replication, the monkeypox and other orthopox viruses have evolved mechanisms to evade host immune cells.[15]


Replication and life cycle
This section may be too technical for most readers to understand. (June 2023) |
As an Orthopoxvirus, MPV replication occurs entirely in the cell cytoplasm within 'factories' – created from the host rough endoplasmic reticulum (ER) – where viral mRNA transcription and translation also take place.[16][17] The factories are also where DNA replication, gene expression, and mature virions (MV) are created.[18]
MVs are able to bind to the cell surface with the help of viral proteins.[19] Virus entry into the host cell plasma membrane is dependent on a neutral pH, otherwise entry occurs via a low-pH dependent endocytic route.[19] The MV of the monkeypox virus has an Entry Fusion Complex (EFC), allowing it to enter the host cell after attachment.[19]
Translation of mRNA into structural virions occurs using the host ribosomes.[16] Gene expression begins when MPV releases viral proteins and enzymatic factors that disable the cell.[20] Mature virions are infectious, however, they will stay inside the cell, until they are transported from the factories to the Golgi/endosomal comportment.[18] Protein synthesis allows for the ER membrane of the factory to dismantle, while small two lipid bilayer membranes will appear to encapsulate the genomes of new virions, now extracellular viruses (EVs).[20][16][18] The VPS52 and VPS54 genes of the GARP complex, which is important for transport, are necessary for wrapping the virus, and formation of EVs.[18] DNA concatemers process the genomes, which appear in new virions, along with other enzymes, and genetic information needed for the replication cycle to occur.[20] EVs are necessary for the spread of the virus from cell-to-cell and its long-distance spread.[18]
Transmission
Animal to human
Zoonotic transmission can occur from direct contact with the blood, bodily fluids, wounds, or mucosal lesions of infected animals whether they are dead or alive. The virus is thought to have originated in Africa where evidence of the virus has been observed in multiple animals including rope squirrels, tree squirrels, Gambian pouched rats, dormice, and different species of monkeys. Though the natural reservoir of the monkeypox virus has not yet been established, rodents are speculated to be the most likely reservoir. Eating meat that has not been properly cooked and consuming other products of infected animals proves to be a major risk factor in the spread of infection.[21]
Human to human

Monkeypox virus can be transmitted from one person to another through contact with infectious lesion material or fluid on the skin, in the mouth or on the genitals; this includes touching, close contact and during sex. It may also spread by means of respiratory droplets from talking, coughing or sneezing.[4][23] During the 2022-2023 outbreak, transmission between people was almost exclusively via sexual contact.[24] There is a lower risk of infection from fomites (objects which can become infectious after being touched by an infected person) such as clothing or bedding, but precautions should be taken.[4]
The virus then enters the body through broken skin, or mucosal surfaces such as the mouth, respiratory tract, or genitals.[25][26]
Human to animal
There are two recorded instances of human to animal transmission. Both occurred during the 2022–2023 global mpox outbreak. In both cases, the owners of a pet dog first became infected with mpox and transmitted the infection to the pet.[27][26]
Mpox disease
Human
Initial symptoms of mpox infection are fever, muscle pains, and sore throat, followed by an itchy or painful rash, headache, swollen lymph nodes, and fatigue. Not everyone will exhibit the complete range of symptoms.[28][29]
Most mpox patients become symptomatic 4–11 days after infection. However, the incubation period can be as short as 1 day. The 2022–2023 outbreak revealed that incubation periods of up to 4 weeks are possible, with 5% of cases having incubation periods longer than the previously assumed 21 days.[28][30]Animal
Prevention
The MVA-BN vaccine, originally developed for smallpox, has been approved for use by persons who are either considered at high risk of exposure to mpox, or who may have recently been exposed to it.[33][34][35] The United States Centers for Disease Control and Prevention (CDC) recommends that persons investigating mpox outbreaks, those caring for infected individuals or animals, and those exposed by close or intimate contact with infected individuals or animals should receive a vaccination.[36]
The CDC has made detailed recommendations in addition to the standard precautions for infection control. These include that healthcare providers don a gown, mask, goggles, and a disposable filtering respirator (such as an N95), and that an infected person should be isolated a private room to keep others from possible contact.[37]Treatment
Immune system interaction
Pox viruses have mechanisms to evade the hosts' innate and adaptive immune systems. Viral proteins, expressed by infected cells, employ multiple approaches to limit immune system activity; including binding to, and preventing activation of proteins within the host's immune system, and preventing infected cells from dying to enable them to continue replicating the monkey pox virus.[44]
Variants and clades
The virus is subclassified into two clades, Clade I and Clade II.[4] At the protein level, the clades share 170 orthologs, and their transcriptional regulatory sequences show no significant differences.[8] Both clades have 53 common virulence genes, which contain different types of amino acid changes. 121 of the amino acid changes in the virulence genes are silent, while 61 are conservative, and 93 are non-conservative.[8]
Historically, the case fatality rate (CFR) of past outbreaks was estimated at between 1% and 10%, with Clade I considered to be more severe than Clade II.[45] The CFR of the 2022-2023 global outbreak (caused by Clade IIb) has been very low - estimated at 0.16%, with the majority of deaths in individuals who were already immunocompromised.[46]
| Name[47] | Former names[47] | Nations[48][49] | |
|---|---|---|---|
| Clade I | Congo Basin
Central African |
||
| Clade II | Clade IIa | West African | |
| Clade IIb | Widespread globally - See 2022–2023 mpox outbreak § Cases per country and territory | ||
History
Monkeypox virus was first identified by Preben von Magnus in Copenhagen, Denmark, in 1958 in crab-eating macaque monkeys (Macaca fascicularis) being used as laboratory animals.[50] The virus was originally given the name monkeypox virus because it had been isolated from monkeys; subsequent research reveals that monkeys are not the main host. Other small mammals in the tropical forests of Central and West Africa.[51] are suspected to form a natural reservoir.[8]
The first human infection was diagnosed 1970, in the Democratic Republic of Congo.[4] Small viral outbreaks with secondary human-to-human infection occur routinely in endemic areas of Central Africa; the primary route of infection is thought to be contact with the infected animals or their bodily fluids.[52] The first reported outbreak in humans outside of Africa occurred in 2003 in the United States; it was traced to Gambian pouched rats which had been imported as exotic pets.[53] There have subsequently been a number of outbreaks to regions outside of the endemic areas in Central Africa.
Notes
- ^ The World Health Organization (the authority on disease names) announced the new name "mpox" in November 2022. But virus naming is the responsibility of the International Committee on the Taxonomy of Viruses (ICTV), which is currently reviewing all orthopoxvirus species. As of March 2023, the official name of the virus remains "monkeypox virus".[2]
References
- ^ "Mpox (monkeypox) outbreak 2022". www.who.int. Archived from the original on 2023-01-07. Retrieved 2023-01-07.
- ^ "WHO recommends new name for monkeypox disease" (Press release). World Health Organization (WHO). 28 November 2022. Retrieved 29 November 2022.
- ^ Hibbert CM (2022-08-11). "Baby boomer alert: Will your childhood smallpox vaccine protect against monkeypox?". News @ Northeastern. Archived from the original on 2022-11-18. Retrieved 2022-11-18.
- ^ a b c d e f g h "WHO Factsheet – Mpox (Monkeypox)". World Health Organization (WHO). 18 April 2023. Retrieved 21 May 2023.
- ^ CDC (2022-10-18). "Monkeypox in the U.S." Centers for Disease Control and Prevention. Archived from the original on 2022-10-26. Retrieved 2022-10-26.
- ^ Bunge, Eveline M.; Hoet, Bernard; Chen, Liddy; Lienert, Florian; Weidenthaler, Heinz; Baer, Lorraine R.; Steffen, Robert (11 February 2022). "The changing epidemiology of human monkeypox – A potential threat? A systematic review". PLOS Neglected Tropical Diseases. 16 (2): e0010141. doi:10.1371/journal.pntd.0010141. PMC 8870502. PMID 35148313.
- ^ "Mpox in Animals | Mpox | Poxvirus | CDC". www.cdc.gov. 2023-04-27. Retrieved 2023-06-08.
- ^ a b c d Lansiaux E, Jain N, Laivacuma S, Reinis A (December 2022). "The virology of human monkeypox virus (hMPXV): A brief overview". Virus Research. 322: 198932. doi:10.1016/j.virusres.2022.198932. PMC 9534104. PMID 36165924.
- ^ a b Shchelkunov SN, Totmenin AV, Babkin IV, Safronov PF, Ryazankina OI, Petrov NA, et al. (November 2001). "Human monkeypox and smallpox viruses: genomic comparison". FEBS Letters. 509 (1): 66–70. doi:10.1016/S0014-5793(01)03144-1. PMC 9533818. PMID 11734207.
- ^ Alakunle E, Moens U, Nchinda G, Okeke MI (November 2020). "Monkeypox Virus in Nigeria: Infection Biology, Epidemiology, and Evolution". Viruses. 12 (11): 1257. doi:10.3390/v12111257. PMC 7694534. PMID 33167496.
- ^ a b c Kaler J, Hussain A, Flores G, Kheiri S, Desrosiers D (July 2022). "Monkeypox: A Comprehensive Review of Transmission, Pathogenesis, and Manifestation". Cureus. 14 (7): e26531. doi:10.7759/cureus.26531. PMC 9345383. PMID 35928395.
- ^ Forni, Diego; Cagliani, Rachele; Molteni, Cristian; Clerici, Mario; Sironi, Manuela (November 2022). "Monkeypox virus: The changing facets of a zoonotic pathogen". Infection, Genetics and Evolution. 105: 105372. doi:10.1016/j.meegid.2022.105372. PMC 9534092. PMID 36202208.
- ^ Kugelman JR, Johnston SC, Mulembakani PM, Kisalu N, Lee MS, Koroleva G, et al. (February 2014). "Genomic variability of monkeypox virus among humans, Democratic Republic of the Congo". Emerging Infectious Diseases. 20 (2): 232–239. doi:10.3201/eid2002.130118. PMC 3901482. PMID 24457084.
- ^ "Monkeypox: What We Do and Don't Know About Recent Outbreaks". ASM.org. Archived from the original on 2022-10-21. Retrieved 2022-10-21.
- ^ Zandi, Milad; Shafaati, Maryam; Hosseini, Fatemeh (2023-02-01). "Mechanisms of immune evasion of monkeypox virus". Frontiers in Microbiology. 14. doi:10.3389/fmicb.2023.1106247. ISSN 1664-302X. PMC 9928851. PMID 36819041.
- ^ a b c "Monkeypox: What We Do and Don't Know About Recent Outbreaks". ASM.org. Archived from the original on 2022-10-21. Retrieved 2022-10-26.
- ^ Moss B (September 2013). "Poxvirus DNA replication". Cold Spring Harbor Perspectives in Biology. 5 (9): a010199. doi:10.1101/cshperspect.a010199. PMC 3753712. PMID 23838441.
- ^ a b c d e Realegeno S, Puschnik AS, Kumar A, Goldsmith C, Burgado J, Sambhara S, et al. (June 2017). "Monkeypox Virus Host Factor Screen Using Haploid Cells Identifies Essential Role of GARP Complex in Extracellular Virus Formation". Journal of Virology. 91 (11): e00011–17. doi:10.1128/JVI.00011-17. PMC 5432867. PMID 28331092.
- ^ a b c Moss B (December 2016). "Membrane fusion during poxvirus entry". Seminars in Cell & Developmental Biology. 60: 89–96. doi:10.1016/j.semcdb.2016.07.015. PMC 5161597. PMID 27423915.
- ^ a b c Alkhalil A, Hammamieh R, Hardick J, Ichou MA, Jett M, Ibrahim S (July 2010). "Gene expression profiling of monkeypox virus-infected cells reveals novel interfaces for host-virus interactions". Virology Journal. 7: 173. doi:10.1186/1743-422X-7-173. PMC 2920256. PMID 20667104.
- ^ "Monkeypox". www.who.int. Archived from the original on 2022-04-21. Retrieved 2022-11-18.
- ^ Kaler, Jasndeep; Hussain, Azhar; Flores, Gina; Kheiri, Shehreen; Desrosiers, Dara (2022). "Monkeypox: A Comprehensive Review of Transmission, Pathogenesis, and Manifestation". Cureus. 14 (7): e26531. doi:10.7759/cureus.26531. ISSN 2168-8184. PMC 9345383. PMID 35928395.
- ^ "Mpox - How It Spreads". Centers for Disease Control and Prevention. 2 February 2023. Archived from the original on 21 May 2023. Retrieved 23 May 2023.
- ^ "Safer Sex, Social Gatherings, and Mpox". Centers for Disease Control and Prevention. 28 April 2023. Archived from the original on 29 May 2023. Retrieved 26 May 2023.
- ^ "WHO Factsheet – Mpox (Monkeypox)". World Health Organization (WHO). 18 April 2023. Archived from the original on 21 April 2022. Retrieved 21 May 2023.
- ^ a b "Brazil: Domestic puppy in Minas Gerais contracts monkeypox, Lived with confirmed human case". Outbreak News Today. 2022-08-30. Retrieved 2023-06-10.
- ^ Seang, Sophie; Burrel, Sonia; Todesco, Eve; Leducq, Valentin; Monsel, Gentiane; Le Pluart, Diane; Cordevant, Christophe; Pourcher, Valérie; Palich, Romain (August 2022). "Evidence of human-to-dog transmission of monkeypox virus". The Lancet. 400 (10353): 658–659. doi:10.1016/s0140-6736(22)01487-8. ISSN 0140-6736. PMC 9536767. PMID 35963267.
- ^ a b c "WHO Factsheet – Mpox (Monkeypox)". World Health Organization (WHO). 18 April 2023. Archived from the original on 21 April 2022. Retrieved 21 May 2023.
- ^ "Mpox Symptoms". U.S. Centers for Disease Control and Prevention (CDC). 2 February 2023. Archived from the original on 21 May 2023. Retrieved 21 May 2023.
- ^ McFarland SE, Marcus U, Hemmers L, Miura F, Iñigo Martínez J, Martínez FM, Montalbán EG, Chazelle E, Mailles A, Silue Y, Hammami N, Lecompte A, Ledent N, Vanden Berghe W, Liesenborghs L, Van den Bossche D, Cleary PR, Wallinga J, Robinson EP, Johansen TB, Bormane A, Melillo T, Seidl C, Coyer L, Boberg R, Jurke A, Werber D, Bartel A (July 2023). "Estimated incubation period distributions of mpox using cases from two international European festivals and outbreaks in a club in Berlin, May to June 2022". Euro Surveillance. 28 (27). doi:10.2807/1560-7917.ES.2023.28.27.2200806. PMC 10370040. PMID 37410383.
- ^ a b "Mpox in Animals". Centers for Disease Control and Prevention. 4 January 2023. Retrieved 25 May 2023.
- ^ Nash SL, Palmer SB, Wingfield WE (2009). "1.11. Zoonoses and zoonotic diseases". In Wingfield WE, Palmer SB (eds.). Veterinary Disaster Response. Iowa: John Wiley & Sons. pp. 167–168. ISBN 978-0-8138-1014-0.
- ^ "Jynneos Vaccine Effectiveness". Centers for Disease Control and Prevention. 2023-05-19. Archived from the original on 24 May 2023. Retrieved 2023-05-24.
- ^ "Considerations on posology for the use of the vaccine Jynneos/ Imvanex (MVA-BN) against monkeypox" (PDF). European Medicines Agency. 19 August 2022. Archived (PDF) from the original on 28 May 2023. Retrieved 28 May 2023.
- ^ "Protecting you from mpox (monkeypox): information on the smallpox vaccination". GOV.UK. Archived from the original on 28 May 2023. Retrieved 2023-05-28.
- ^ "About Mpox". U.S. Centers for Disease Control and Prevention (CDC). Archived from the original on 11 March 2023. Retrieved 13 March 2023.
- ^ "Infection Prevention and Control of Mpox in Healthcare Settings". U.S. Centers for Disease Control and Prevention (CDC). 31 October 2022. Archived from the original on 18 May 2022. Retrieved 21 May 2022.
- ^ a b "Mpox (monkeypox)". World Health Organization. 12 May 2023. Archived from the original on 23 May 2023. Retrieved 24 May 2023.
- ^ "Patient's Guide to Mpox Treatment with Tecovirimat (TPOXX)". Centers for Disease Control and Prevention. 28 November 2022. Archived from the original on 24 May 2023. Retrieved 2023-05-24.
- ^ "Tecovirimat SIGA". European Medicines Agency. 28 January 2022. Archived from the original on 16 May 2022. Retrieved 19 May 2022.
- ^ "Mpox (formerly Monkeypox)". NIH: National Institute of Allergy and Infectious Diseases. 6 December 2022. Archived from the original on 23 May 2023. Retrieved 2023-05-24.
- ^ a b Fox T, Gould S, Princy N, Rowland T, Lutje V, Kuehn R (March 2023). Cochrane Infectious Diseases Group (ed.). "Therapeutics for treating mpox in humans". The Cochrane Database of Systematic Reviews. 2023 (3): CD015769. doi:10.1002/14651858.CD015769. PMC 10012405. PMID 36916727.
- ^ "Mpox (monkeypox) – Treatment algorithm". BMJ Best Practice. Archived from the original on 4 December 2022. Retrieved 14 March 2023.
- ^ Lum, Fok-Moon; Torres-Ruesta, Anthony; Tay, Matthew Z.; Lin, Raymond T. P.; Lye, David C.; Rénia, Laurent; Ng, Lisa F. P. (September 2022). "Monkeypox: disease epidemiology, host immunity and clinical interventions". Nature Reviews Immunology. 22 (10): 597–613. doi:10.1038/s41577-022-00775-4. ISSN 1474-1741. PMC 9443635. PMID 36064780.
- ^ Vogel, Lauren (2022-08-15). "Making sense of monkeypox death rates". CMAJ. 194 (31): E1097. doi:10.1503/cmaj.1096012. ISSN 0820-3946. PMC 9377567. PMID 35970550. Archived from the original on 31 May 2023. Retrieved 31 May 2023.
- ^ "Mpox (monkeypox) - Prognosis". BMJ Best Practice. Archived from the original on 31 May 2023. Retrieved 2023-05-31.
- ^ a b "Monkeypox: experts give virus variants new names". www.who.int. Archived from the original on 2022-08-19. Retrieved 2022-08-19.
- ^ "Monkeypox". www.who.int. Archived from the original on 2022-04-21. Retrieved 2022-08-19.
- ^ Likos AM, Sammons SA, Olson VA, Frace AM, Li Y, Olsen-Rasmussen M, et al. (October 2005). "A tale of two clades: monkeypox viruses". The Journal of General Virology. 86 (Pt 10): 2661–2672. doi:10.1099/vir.0.81215-0. PMID 16186219.
- ^ "Monkeypox". New Scientist. 80. Reed Business Information: 682–. 30 November 1978. ISSN 0262-4079. Archived from the original on 13 January 2023. Retrieved 3 November 2016.
- ^ Igiebor FA, Agbontaen OJ, Egharevba PA, Amengialue OO, Ehiaghe JI, Ovwero E, Ehiaghe FA (May 2022). "Monkeypox: Emerging and Re-Emerging Threats in Nigeria". Journal of Basic and Applied Sciences. 7 (1). Benin City, Nigeria: Faculty of Science, Benson Idahosa University: 119–132. Archived from the original on 2022-06-07. Retrieved 2022-06-07.
- ^ Meyer H, Perrichot M, Stemmler M, Emmerich P, Schmitz H, Varaine F, et al. (August 2002). "Outbreaks of disease suspected of being due to human monkeypox virus infection in the Democratic Republic of Congo in 2001". Journal of Clinical Microbiology. 40 (8): 2919–2921. doi:10.1128/JCM.40.8.2919-2921.2002. PMC 120683. PMID 12149352.
- ^ "2003 U.S. Outbreak Monkeypox". CDC. 11 May 2015. Archived from the original on 15 October 2017. Retrieved 15 October 2017.
External links
- CDC Questions and Answers About Monkeypox
- CDC – Human Monkeypox – Kasai Oriental, Zaire, 1996–1997 Archived 2022-05-23 at the Wayback Machine
- CDC – Outbreak of Human Monkeypox, Democratic Republic of Congo, 1996 to 1997
- CDC Preliminary Report: Multistate Outbreak of Monkeypox in Persons Exposed to Pet Prairie Dogs
- National Library of Medicine – Monkeypox virus
- Virology.net Picturebook: Monkeypox Archived 2005-10-28 at the Wayback Machine
- Viralzone: Orthopoxvirus
- Virus Pathogen Database and Analysis Resource (ViPR): Poxviridae
- World Health Organization to rename the Monkeypox Virus
