Pseudomonas
| Pseudomonas | |
|---|---|
| |
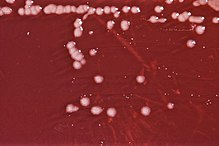
| P. aeruginosa colonies on an agar plate | |
| Scientific classification | |
| Domain: | Bacteria |
| Phylum: | Pseudomonadota |
| Class: | Gammaproteobacteria |
| Order: | Pseudomonadales |
| Family: | Pseudomonadaceae |
| Genus: | Pseudomonas Migula 1894 |
| Type species | |
| Pseudomonas aeruginosa | |
| Species | |
|
See text. | |
| Synonyms | |
| |
Pseudomonas is a genus of Gram-negative bacteria belonging to the family Pseudomonadaceae in the class Gammaproteobacteria. The 313 members of the genus[2][3] demonstrate a great deal of metabolic diversity and consequently are able to colonize a wide range of niches.[4] Their ease of culture in vitro and availability of an increasing number of Pseudomonas strain genome sequences has made the genus an excellent focus for scientific research; the best studied species include P. aeruginosa in its role as an opportunistic human pathogen, the plant pathogen P. syringae, the soil bacterium P. putida, and the plant growth-promoting P. fluorescens, P. lini, P. migulae, and P. graminis.[5][6]
Because of their widespread occurrence in water and plant seeds such as dicots, the pseudomonads were observed early in the history of microbiology. The generic name Pseudomonas created for these organisms was defined in rather vague terms by Walter Migula in 1894 and 1900 as a genus of Gram-negative, rod-shaped, and polar-flagellated bacteria with some sporulating species.[7][8] The latter statement was later proved incorrect and was due to refractive granules of reserve materials.[9] Despite the vague description, the type species, Pseudomonas pyocyanea (basonym of Pseudomonas aeruginosa), proved the best descriptor.[9]
Classification history[edit]
Like most bacterial genera, the pseudomonad[note 1] last common ancestor lived hundreds of millions of years ago. They were initially classified at the end of the 19th century when first identified by Walter Migula. The etymology of the name was not specified at the time and first appeared in the seventh edition of Bergey's Manual of Systematic Bacteriology (the main authority in bacterial nomenclature) as Greek pseudes (ψευδής) "false" and -monas (μονάς/μονάδος) "a single unit", which can mean false unit; however, Migula possibly intended it as false Monas, a nanoflagellated protist[9] (subsequently, the term "monad" was used in the early history of microbiology to denote unicellular organisms). Soon, other species matching Migula's somewhat vague original description were isolated from many natural niches and, at the time, many were assigned to the genus. However, many strains have since been reclassified, based on more recent methodology and use of approaches involving studies of conservative macromolecules.[10]
Recently, 16S rRNA sequence analysis has redefined the taxonomy of many bacterial species.[11] As a result, the genus Pseudomonas includes strains formerly classified in the genera Chryseomonas and Flavimonas.[12] Other strains previously classified in the genus Pseudomonas are now classified in the genera Burkholderia and Ralstonia.[13][14]
In 2020, a phylogenomic analysis of 494 complete Pseudomonas genomes identified two well-defined species (P. aeruginosa and P. chlororaphis) and four wider phylogenetic groups (P. fluorescens, P. stutzeri, P. syringae, P. putida) with a sufficient number of available proteomes.[15] The four wider evolutionary groups include more than one species, based on species definition by the Average Nucleotide Identity levels.[16] In addition, the phylogenomic analysis identified several strains that were mis-annotated to the wrong species or evolutionary group.[15] This mis-annotation problem has been reported by other analyses as well.[17]
Genomics[edit]
In 2000, the complete genome sequence of a Pseudomonas species was determined; more recently, the sequence of other strains has been determined, including P. aeruginosa strains PAO1 (2000), P. putida KT2440 (2002), P. protegens Pf-5 (2005), P. syringae pathovar tomato DC3000 (2003), P. syringae pathovar syringae B728a (2005), P. syringae pathovar phaseolica 1448A (2005), P. fluorescens Pf0-1, and P. entomophila L48.[10]
By 2016, more than 400 strains of Pseudomonas had been sequenced.[18] Sequencing the genomes of hundreds of strains revealed highly divergent species within the genus. In fact, many genomes of Pseudomonas share only 50-60% of their genes, e.g. P. aeruginosa and P. putida share only 2971 proteins out of 5350 (or ~55%).[18]
By 2020, more than 500 complete Pseudomonas genomes were available in Genebank. A phylogenomic analysis utilized 494 complete proteomes and identified 297 core orthologues, shared by all strains.[15] This set of core orthologues at the genus level was enriched for proteins involved in metabolism, translation, and transcription and was utilized for generating a phylogenomic tree of the entire genus, to delineate the relationships among the Pseudomonas major evolutionary groups.[15] In addition, group-specific core proteins were identified for most evolutionary groups, meaning that they were present in all members of the specific group, but absent in other pseudomonads. For example, several P. aeruginosa-specific core proteins were identified that are known to play an important role in this species' pathogenicity, such as CntL, CntM, PlcB, Acp1, MucE, SrfA, Tse1, Tsi2, Tse3, and EsrC.[15]
Characteristics[edit]
Members of the genus display these defining characteristics:[19]
- Rod-shaped
- Gram-negative
- Flagellum one or more, providing motility
- Aerobic
- Non-spore forming
- Catalase-positive
- Oxidase-variable
Other characteristics that tend to be associated with Pseudomonas species (with some exceptions) include secretion of pyoverdine, a fluorescent yellow-green siderophore[20] under iron-limiting conditions. Certain Pseudomonas species may also produce additional types of siderophore, such as pyocyanin by Pseudomonas aeruginosa[21] and thioquinolobactin by Pseudomonas fluorescens.[22] Pseudomonas species also typically give a positive result to the oxidase test, the absence of gas formation from glucose, glucose is oxidised in oxidation/fermentation test using Hugh and Leifson O/F test, beta hemolytic (on blood agar), indole negative, methyl red negative, Voges–Proskauer test negative, and citrate positive.[citation needed]
Pseudomonas may be the most common nucleator of ice crystals in clouds, thereby being of utmost importance to the formation of snow and rain around the world.[23]
Biofilm formation[edit]
All species and strains of Pseudomonas have historically been classified as strict aerobes. Exceptions to this classification have recently been discovered in Pseudomonas biofilms.[24] A significant number of cells can produce exopolysaccharides associated with biofilm formation. Secretion of exopolysaccharides such as alginate makes it difficult for pseudomonads to be phagocytosed by mammalian white blood cells.[25] Exopolysaccharide production also contributes to surface-colonising biofilms that are difficult to remove from food preparation surfaces. Growth of pseudomonads on spoiling foods can generate a "fruity" odor.[citation needed]
Antibiotic resistance[edit]
Most Pseudomonas spp. are naturally resistant to penicillin and the majority of related beta-lactam antibiotics, but a number are sensitive to piperacillin, imipenem, ticarcillin, or ciprofloxacin.[25] Aminoglycosides such as tobramycin, gentamicin, and amikacin are other choices for therapy.[citation needed]
This ability to thrive in harsh conditions is a result of their hardy cell walls that contain porins. Their resistance to most antibiotics is attributed to efflux pumps, which pump out some antibiotics before they are able to act.[citation needed]
Pseudomonas aeruginosa is increasingly recognized as an emerging opportunistic pathogen of clinical relevance. One of its most worrying characteristics is its low antibiotic susceptibility.[26] This low susceptibility is attributable to a concerted action of multidrug efflux pumps with chromosomally encoded antibiotic resistance genes (e.g., mexAB-oprM, mexXY, etc.[27]) and the low permeability of the bacterial cellular envelopes. Besides intrinsic resistance, P. aeruginosa easily develops acquired resistance either by mutation in chromosomally encoded genes or by the horizontal gene transfer of antibiotic resistance determinants. Development of multidrug resistance by P. aeruginosa isolates requires several different genetic events that include acquisition of different mutations and/or horizontal transfer of antibiotic resistance genes. Hypermutation favours the selection of mutation-driven antibiotic resistance in P. aeruginosa strains producing chronic infections, whereas the clustering of several different antibiotic resistance genes in integrons favours the concerted acquisition of antibiotic resistance determinants. Some recent studies have shown phenotypic resistance associated to biofilm formation or to the emergence of small-colony-variants, which may be important in the response of P. aeruginosa populations to antibiotic treatment.[10]
Sensitivity to gallium[edit]
Although gallium has no natural function in biology, gallium ions interact with cellular processes in a manner similar to iron(III). When gallium ions are mistakenly taken up in place of iron(III) by bacteria such as Pseudomonas, the ions interfere with respiration, and the bacteria die. This happens because iron is redox-active, allowing the transfer of electrons during respiration, while gallium is redox-inactive.[28][29]
Pathogenicity[edit]
Animal pathogens[edit]
Infectious species include P. aeruginosa, P. oryzihabitans, and P. plecoglossicida. P. aeruginosa flourishes in hospital environments, and is a particular problem in this environment, since it is the second-most common infection in hospitalized patients (nosocomial infections).[30] This pathogenesis may in part be due to the proteins secreted by P. aeruginosa. The bacterium possesses a wide range of secretion systems, which export numerous proteins relevant to the pathogenesis of clinical strains.[31] Intriguingly, several genes involved in the pathogenesis of P. aeruginosa, such as CntL, CntM, PlcB, Acp1, MucE, SrfA, Tse1, Tsi2, Tse3, and EsrC are core group-specific,[15] meaning that they are shared by the vast majority of P. aeruginosa strains, but they are not present in other Pseudomonads.
Plant pathogens[edit]
P. syringae is a prolific plant pathogen. It exists as over 50 different pathovars, many of which demonstrate a high degree of host-plant specificity. Numerous other Pseudomonas species can act as plant pathogens, notably all of the other members of the P. syringae subgroup, but P. syringae is the most widespread and best-studied.[citation needed]
Fungus pathogens[edit]
P. tolaasii can be a major agricultural problem, as it can cause bacterial blotch of cultivated mushrooms.[32] Similarly, P. agarici can cause drippy gill in cultivated mushrooms.[33]
Use as biocontrol agents[edit]
Since the mid-1980s, certain members of the genus Pseudomonas have been applied to cereal seeds or applied directly to soils as a way of preventing the growth or establishment of crop pathogens. This practice is generically referred to as biocontrol. The biocontrol properties of P. fluorescens and P. protegens strains (CHA0 or Pf-5 for example) are currently best-understood, although it is not clear exactly how the plant growth-promoting properties of P. fluorescens are achieved. Theories include: the bacteria might induce systemic resistance in the host plant, so it can better resist attack by a true pathogen; the bacteria might outcompete other (pathogenic) soil microbes, e.g. by siderophores giving a competitive advantage at scavenging for iron; the bacteria might produce compounds antagonistic to other soil microbes, such as phenazine-type antibiotics or hydrogen cyanide. Experimental evidence supports all of these theories.[34]
Other notable Pseudomonas species with biocontrol properties include P. chlororaphis, which produces a phenazine-type antibiotic active agent against certain fungal plant pathogens,[35] and the closely related species P. aurantiaca, which produces di-2,4-diacetylfluoroglucylmethane, a compound antibiotically active against Gram-positive organisms.[36]
Use as bioremediation agents[edit]
Some members of the genus are able to metabolise chemical pollutants in the environment, and as a result, can be used for bioremediation. Notable species demonstrated as suitable for use as bioremediation agents include:
- P. alcaligenes, which can degrade polycyclic aromatic hydrocarbons.[37]
- P. mendocina, which is able to degrade toluene.[38]
- P. pseudoalcaligenes, which is able to use cyanide as a nitrogen source.[39]
- P. resinovorans, which can degrade carbazole.[40]
- P. aeruginosa, P. putida, P. desmolyticum, and P. nitroreducens can degrade chlorpyrifos.[41]
- P. veronii, which has been shown to degrade a variety of simple aromatic organic compounds.[42][43]
- P. putida, which has the ability to degrade organic solvents such as toluene.[44] At least one strain of this bacterium is able to convert morphine in aqueous solution into the stronger and somewhat expensive to manufacture drug hydromorphone (Dilaudid).
- Strain KC of P. stutzeri, which is able to degrade carbon tetrachloride.[45]
Risks associated with pseudomonas[edit]
Pseudomonas is a genus of bacteria known to be associated with several diseases affecting humans, plants, and animals.
Humans & Animals[edit]
One of the most concerning strains of Pseudomonas is Pseudomonas aeruginosa, which is responsible for a considerable number of hospital-acquired infections. Numerous hospitals and medical facilities face persistent challenges in dealing with Pseudomonas infections. The symptoms of these infections are caused by proteins secreted by the bacteria and may include pneumonia, blood poisoning, and urinary tract infections.[46] Pseudomonas aeruginosa is highly contagious and has displayed resistance to antibiotic treatments, making it difficult to manage effectively. Some strains of Pseudomonas are known to target white blood cells in various mammal species, posing risks to humans, cattle, sheep, and dogs alike.[47]
Fish[edit]
While Pseudomonas aeruginosa seems to be a pathogen that primarily affects humans, another strain known as Pseudomonas plecoglossicida poses risks to fish. This strain can cause gastric swelling and haemorrhaging in fish populations.[47]
Plants & Fungi[edit]
Various strains of Pseudomonas are recognized as pathogens in the plant kingdom. Notably, the Pseudomonas syringae family is linked to diseases affecting a wide range of agricultural plants, with different strains showing adaptations to specific host species. In particular, the virulent strain Pseudomonas tolaasii is responsible for causing blight and degradation in edible mushroom species.[47]
Detection of food spoilage agents in milk[edit]
One way of identifying and categorizing multiple bacterial organisms in a sample is to use ribotyping.[48] In ribotyping, differing lengths of chromosomal DNA are isolated from samples containing bacterial species, and digested into fragments.[48] Similar types of fragments from differing organisms are visualized and their lengths compared to each other by Southern blotting or by the much faster method of polymerase chain reaction (PCR).[48] Fragments can then be matched with sequences found on bacterial species.[48] Ribotyping is shown to be a method to isolate bacteria capable of spoilage.[49] Around 51% of Pseudomonas bacteria found in dairy processing plants are P. fluorescens, with 69% of these isolates possessing proteases, lipases, and lecithinases which contribute to degradation of milk components and subsequent spoilage.[49] Other Pseudomonas species can possess any one of the proteases, lipases, or lecithinases, or none at all.[49] Similar enzymatic activity is performed by Pseudomonas of the same ribotype, with each ribotype showing various degrees of milk spoilage and effects on flavour.[49] The number of bacteria affects the intensity of spoilage, with non-enzymatic Pseudomonas species contributing to spoilage in high number.[49]
Food spoilage is detrimental to the food industry due to production of volatile compounds from organisms metabolizing the various nutrients found in the food product.[50] Contamination results in health hazards from toxic compound production as well as unpleasant odours and flavours.[50] Electronic nose technology allows fast and continuous measurement of microbial food spoilage by sensing odours produced by these volatile compounds.[50] Electronic nose technology can thus be applied to detect traces of Pseudomonas milk spoilage and isolate the responsible Pseudomonas species.[51] The gas sensor consists of a nose portion made of 14 modifiable polymer sensors that can detect specific milk degradation products produced by microorganisms.[51] Sensor data is produced by changes in electric resistance of the 14 polymers when in contact with its target compound, while four sensor parameters can be adjusted to further specify the response.[51] The responses can then be pre-processed by a neural network which can then differentiate between milk spoilage microorganisms such as P. fluorescens and P. aureofaciens.[51]
Species[edit]
Pseudomonas comprises the following species,[52] organized into genomic affinity groups:[53][54][55][56][57][58][59]
P. aeruginosa Group[edit]
- P. aeruginosa (Schroeter 1872) Migula 1900 (Approved Lists 1980)
- P. citronellolis Seubert 1960 (Approved Lists 1980)
- P. delhiensis Prakash et al. 2007
- P. denitrificans Bergey et al. 1961
- P. jinjuensis Kwon et al. 2003
- P. knackmussii Stolz et al. 2007
- P. nicosulfuronedens Li et al. 2021
- P. nitroreducens Iizuka and Komagata 1964 (Approved Lists 1980)
- P. panipatensis Gupta et al. 2008
P. anguilliseptica Group[edit]
- P. anguilliseptica Wakabayashi and Egusa 1972 (Approved Lists 1980)
- P. benzenivorans Lang et al. 2012
- P. borbori Vanparys et al. 2006
- P. campi Timsy et al. 2021
- P. cuatrocienegasensis Escalante et al. 2009
- P. glareae Romanenko et al. 2015
- P. guineae Bozal et al. 2007
- P. guryensis Kim et al. 2021
- P. lalucatii Busquets et al. 2021
- P. leptonychotis Nováková et al. 2020
- P. marincola Romanenko et al. 2008
- P. peli Vanparys et al. 2006
- P. segitis Park et al. 2006
- P. taeanensis Lee et al. 2010
- P. ullengensis Kim et al. 2021
P. fluorescens Group[edit]
P. asplenii Subgroup
- P. agarici Young 1970 (Approved Lists 1980)
- P. asplenii (Ark and Tompkins 1946) Savulescu 1947 (Approved Lists 1980)
- P. batumici Kiprianova et al. 2011
- P. fuscovaginae (ex Tanii et al. 1976) Miyajima et al. 1983
- P. gingeri Cutri et al. 1984
- P. vanderleydeniana Girard et al. 2022
P. chlororaphis Subgroup
- P. aurantiaca Nakhimovskaya 1948 (Approved Lists 1980)
- P. aureofaciens Kluyver 1956 (Approved Lists 1980)
- P. chlororaphis (Guignard and Sauvageau 1894) Bergey et al. 1930 (Approved Lists 1980)
- P. piscium (Burr et al. 2010) Chen et al. 2018
P. corrugata Subgroup
- P. alvandae Girard et al. 2022
- P. bijieensis Liang et al. 2021
- P. brassicacearum Achouak et al. 2000
- P. canavaninivorans Hauth et al. 2022
- P. corrugata Roberts and Scarlett 1981
- P. kilonensis Sikorski et al. 2001
- P. marvdashtae Girard et al. 2022
- P. mediterranea Catara et al. 2002
- P. ogarae Garrido-Sanz et al. 2022
- P. tehranensis Girard et al. 2022
- P. thivervalensis Achouak et al. 2000
- P. viciae Zhao et al. 2020
- P. zanjanensis Girard et al. 2022
- P. zarinae Girard et al. 2022
P. fluorescens Subgroup
- P. allii Sawada et al. 2021
- P. antarctica Reddy et al. 2004
- P. asgharzadehiana Girard et al. 2022
- P. aylmerensis corrig. Tchagang et al. 2021
- P. azadiae Girard et al. 2022
- P. azotoformans Iizuka and Komagata 1963 (Approved Lists 1980)
- P. canadensis Tambong et al. 2017
- P. carnis Lick et al. 2020
- P. cedrina corrig. Dabboussi et al. 2002
- P. costantinii Munsch et al. 2002
- P. cremoris Hofmann et al. 2021
- P. cyclaminis Sawada et al. 2021
- P. edaphica Ramírez-Bahena et al. 2019
- P. extremaustralis López et al. 2010
- P. extremorientalis Ivanova et al. 2002
- P. fildesensis Pavlov et al. 2020
- P. fluorescens Migula 1895 (Approved Lists 1980)
- P. fulgida Naureen et al. 2005
- P. grimontii Baïda et al. 2002
- P. haemolytica Hofmann et al. 2020
- P. kairouanensis Oueslati et al. 2020
- P. karstica Švec et al. 2020
- P. khavaziana Girard et al. 2022
- P. kitaguniensis Sawada et al. 2020
- P. lactis von Neubeck et al. 2017
- P. lactucae Sawada et al. 2021
- P. libanensis Dabboussi et al. 1999
- P. lurida Behrendt et al. 2007
- P. marginalis (Brown 1918) Stevens 1925 (Approved Lists 1980)
- P. nabeulensis Oueslati et al. 2020
- P. orientalis Dabboussi et al. 2002
- P. palleroniana Gardan et al. 2002
- P. panacis Park et al. 2005
- P. paracarnis Lick et al. 2021
- P. paralactis von Neubeck et al. 2017
- P. pisciculturae Duman et al. 2021
- P. poae Behrendt et al. 2003
- P. rhodesiae Coroler et al. 1997
- P. salmasensis Girard et al. 2022
- P. salomonii Gardan et al. 2002
- P. simiae Vela et al. 2006
- P. sivasensis Duman et al. 2020
- P. spelaei Švec et al. 2020
- P. synxantha (Ehrenberg 1840) Holland 1920 (Approved Lists 1980)
- P. tolaasii Paine 1919 (Approved Lists 1980)
- P. tritici Girard et al. 2022
- P. trivialis Behrendt et al. 2003
- P. veronii Elomari et al. 1996
- P. yamanorum Arnau et al. 2015
P. fragi Subgroup
- P. bubulae Lick et al. 2020
- P. deceptionensis Carrión et al. 2011
- P. endophytica Ramírez-Bahena et al. 2015
- P. fragi (Eichholz 1902) Gruber 1905 (Approved Lists 1980)
- P. helleri von Neubeck et al. 2016
- P. lundensis Molin et al. 1986
- P. paraversuta Lick et al. 2021
- P. psychrophila Yumoto et al. 2002
- P. saxonica Hofmann et al. 2020
- P. taetrolens Haynes 1957 (Approved Lists 1980)
- P. versuta See-Too et al. 2017
- P. weihenstephanensis von Neubeck et al. 2016
P. gessardii Subgroup
- P. brenneri Baïda et al. 2002
- P. gessardii Verhille et al. 1999
- P. meridiana Reddy et al. 2004
- P. mucidolens Levine and Anderson 1932 (Approved Lists 1980)
- P. proteolytica Reddy et al. 2004
- P. shahriarae Girard et al. 2022
P. jessenii Subgroup
- P. azerbaijanoccidentalis corrig. Girard et al. 2022
- P. azerbaijanorientalis corrig. Girard et al. 2022
- P. izuensis Lu et al. 2020
- P. jessenii Verhille et al. 1999
- P. laurylsulfatiphila corrig. Furmanczyk et al. 2019
- P. laurylsulfativorans corrig. Furmanczyk et al. 2019
- P. mohnii Cámara et al. 2007
- P. moorei Cámara et al. 2007
- P. reinekei Cámara et al. 2007
- P. umsongensis Kwon et al. 2003
- P. vancouverensis Mohn et al. 1999
P. koreensis Subgroup
- P. allokribbensis Morimoto et al. 2021
- P. anatoliensis Duman et al. 2021
- P. atacamensis Poblete-Morales et al. 2021
- P. atagonensis corrig. Morimoto et al. 2020
- P. baetica López et al. 2012
- P. bananamidigenes Girard et al. 2021
- P. botevensis Girard et al. 2021
- P. crudilactis Schlusselhuber et al. 2021
- P. ekonensis Girard et al. 2022
- P. glycinae Jia et al. 2021
- P. gozinkensis Morimoto et al. 2021
- P. granadensis Pascual et al. 2015
- P. hamedanensis Girard et al. 2022
- P. helmanticensis Ramírez-Bahena et al. 2014
- P. iranensis Girard et al. 2022
- P. iridis Duman et al. 2021
- P. khorasanensis Girard et al. 2022
- P. koreensis Kwon et al. 2003
- P. kribbensis Chang et al. 2016
- P. monsensis Girard et al. 2022
- P. moraviensis Tvrzová et al. 2006
- P. neuropathica Duman et al. 2021
- P. siliginis Girard et al. 2022
- P. tensinigenes Girard et al. 2022
- P. triticicola Girard et al. 2022
- P. zeae Girard et al. 2022
P. mandelii Subgroup
- P. arsenicoxydans Campos et al. 2011
- P. farris Girard et al. 2022
- P. frederiksbergensis Andersen et al. 2000
- P. gregormendelii Kosina et al. 2016
- P. lini Delorme et al. 2002
- P. mandelii Verhille et al. 1999
- P. migulae Verhille et al. 1999
- P. mucoides Duman et al. 2021
- P. piscicola Duman et al. 2021
- P. prosekii Kosina et al. 2014
- P. silesiensis Kaminski et al. 2018
P. protegens Subgroup
- P. aestus Vasconcellos et al. 2017
- P. piscis Liu et al. 2020
- P. protegens Ramette et al. 2012
- P. saponiphila Lang et al. 2012
- P. sessilinigenes Girard et al. 2021
incertae sedis
- P. blatchfordae Blatchford and Schuster 1980
- P. kielensis Gieschler et al. 2021
P. linyingensis Group[edit]
- P. guangdongensis Yang et al. 2013
- P. linyingensis He et al. 2015
- P. oryzae Yu et al. 2013
- P. sagittaria Lin et al. 2013
P. lutea Group[edit]
- P. abietaniphila Mohn et al. 1999
- P. bohemica Saati-Santamaría et al. 2018
- P. graminis Behrendt et al. 1999
- P. lutea Peix et al. 2004
P. massiliensis Group[edit]
- P. massiliensis Bardet et al. 2018
- P. typographi Peral-Aranega et al. 2021
P. oleovorans Group[edit]
- P. alcaliphila Yumoto et al. 2001
- P. chaetocerotis Girard et al.
- P. chengduensis Tao et al. 2014
- P. composti Gibello et al. 2011
- P. guguanensis Liu et al. 2013
- P. hydrolytica Zhou et al. 2020
- P. indoloxydans Manickam et al. 2008
- P. khazarica Tarhriz et al. 2020
- P. mendocina Palleroni 1970 (Approved Lists 1980)
- P. oleovorans Lee and Chandler 1941 (Approved Lists 1980)
- P. sediminis Behera et al. 2018
- P. sihuiensis Wu et al. 2014
- P. toyotomiensis Hirota et al. 2011
P. oryzihabitans Group[edit]
- P. asuensis Reddy and Garcia-Pichel 2015
- P. duriflava Liu et al. 2008
- P. luteola Kodama et al. 1985
- P. oryzihabitans Kodama et al. 1985
- P. rhizoryzae Wang et al. 2020
P. pohangensis Group[edit]
- P. mangrovi Ye et al. 2019
- P. pohangensis Weon et al. 2006
P. putida Group[edit]
- P. akappageensis corrig. Morimoto et al. 2020
- P. alkylphenolica Mulet et al. 2015
- P. alloputida Keshavarz-Tohid et al. 2020
- P. anuradhapurensis Girard et al. 2022
- P. arcuscaelestis Mulet et al. 2021
- P. asiatica Tohya et al. 2019
- P. brassicae Sawada et al. 2020
- P. capeferrum Berendsen et al. 2015
- P. cremoricolorata Uchino et al. 2002
- P. defluvii Qin et al. 2020
- P. donghuensis Gao et al. 2015
- P. entomophila Mulet et al. 2012
- P. fakonensis Girard et al. 2022
- P. farsensis Girard et al. 2022
- P. fulva Iizuka and Komagata 1963 (Approved Lists 1980)
- P. guariconensis Toro et al. 2013
- P. huaxiensis Qin et al. 2019
- P. hunanensis Gao et al. 2014
- P. hutmensis Xiang et al. 2019
- P. inefficax Keshavarz-Tohid et al. 2019
- P. japonica Pungrasmi et al. 2008
- P. juntendi Tohya et al. 2019
- P. kermanshahensis Girard et al. 2022
- P. kurunegalensis Girard et al. 2022
- P. laurentiana Wright et al. 2019
- P. maumuensis Girard et al. 2022
- P. monteilii Elomari et al. 1997
- P. mosselii Dabboussi et al. 2002
- P. muyukensis Girard et al. 2022
- P. oryzicola Girard et al. 2022
- P. oryziphila Yang et al. 2021
- P. palmensis Gutierrez-Albanchez et al. 2022
- P. parafulva Uchino et al. 2002
- P. peradeniyensis Girard et al. 2022
- P. persica Keshavarz-Tohid et al. 2020
- P. plecoglossicida Nishimori et al. 2000
- P. promysalinigenes Girard et al. 2022
- P. putida (Trevisan 1889) Migula 1895 (Approved Lists 1980)
- P. pyomelaminifaciens Chakraborty et al.
- P. qingdaonensis Wang et al. 2019
- P. reidholzensis Frasson et al. 2017
- P. shirazensis Girard et al. 2022
- P. shirazica Keshavarz-Tohid et al. 2020
- P. sichuanensis Qin et al. 2019
- P. soli Pascual et al. 2015
- P. taiwanensis Wang et al. 2010
- P. tructae Oh et al. 2019
- P. urmiensis Girard et al. 2022
- P. vlassakiae Girard et al. 2021
- P. vranovensis Tvrzová et al. 2006
- P. wadenswilerensis Frasson et al. 2017
- P. wayambapalatensis Girard et al. 2021
- P. xantholysinigenes Girard et al. 2022
- P. xanthosomatis corrig. Girard et al. 2022
P. resinovorans Group[edit]
- P. furukawaii Kimura et al. 2018
- P. lalkuanensis Thorat et al. 2020
- P. mangiferae Anurat et al. 2019
- P. otitidis Clark et al. 2006
- P. resinovorans Delaporte et al. 1961 (Approved Lists 1980)
P. rhizosphaerae Group[edit]
- P. baltica Gieschler et al. 2021
- P. coleopterorum Menéndez et al. 2015
- P. eucalypticola Liu et al. 2021
- P. rhizosphaerae Peix et al. 2003
P. straminea Group[edit]
- P. argentinensis Peix et al. 2005
- P. daroniae Bueno-Gonzalez et al. 2019
- P. dryadis Bueno-Gonzalez et al. 2019
- P. flavescens Hildebrand et al. 1994
- P. punonensis Ramos et al. 2013
- P. seleniipraecipitans corrig. Hunter and Manter 2011
- P. straminea corrig. Iizuka and Komagata 1963 (Approved Lists 1980)
P. stutzeri Group[edit]
- P. azotifigens Hatayama et al. 2005
- P. balearica Bennasar et al. 1996
- P. chloritidismutans Wolterink et al. 2002
- P. kirkiae Bueno-Gonzalez et al. 2020
- P. nitrititolerans Peng et al. 2019
- P. nosocomialis Mulet et al. 2019
- P. saudiphocaensis Azhar et al. 2017
- P. songnenensis Zhang et al. 2015
- P. stutzeri (Lehmann and Neumann 1896) Sijderius 1946 (Approved Lists 1980)
- P. urumqiensis Zou et al. 2019
- P. xanthomarina Romanenko et al. 2005
- P. zhaodongensis Zhang et al. 2015
P. syringae Group[edit]
- P. alliivorans Zhao et al. 2021
- P. amygdali Psallidas and Panagopoulos 1975 (Approved Lists 1980)
- P. asturiensis González et al. 2013
- P. avellanae Janse et al. 1997
- P. cannabina (ex Šutič and Dowson 1959) Gardan et al. 1999
- P. capsici Zhao et al. 2021
- P. caricapapayae Robbs 1956 (Approved Lists 1980)
- P. caspiana Busquets et al. 2017
- P. cerasi Kałuzna et al. 2017
- P. cichorii (Swingle 1925) Stapp 1928 (Approved Lists 1980)
- P. congelans Behrendt et al. 2003
- P. coronafaciens (Elliott 1920) Stevens 1958
- P. ficuserectae Goto 1983
- P. floridensis Timilsina et al. 2018
- P. foliumensis Tambong et al. 2021
- P. helianthi Elasri et al. 2001
- P. meliae Ogimi 1981
- P. ovata Rao et al. 2021
- P. savastanoi (Janse 1982) Gardan et al. 1992
- P. syringae van Hall 1902 (Approved Lists 1980)
- P. tomato Gardan et al. 1999
- P. tremae Gardan et al. 1999
- P. triticumensis Tambong et al. 2021
- P. viridiflava (Burkholder 1930) Dowson 1939 (Approved Lists 1980)
incertae sedis[edit]
- P. acephalitica Tapia-Paniagua et al. 2014
- P. acidophila Imada et al. 1981
- "Ca. P. adelgestsugas" von Dohlen et al. 2013
- P. alcaligenes Monias 1928 (Approved Lists 1980)
- P. alginovora Boyen et al. 1990
- P. alkanolytica Nakao and Kuno 1972
- P. amyloderamosa Norrman and Wober 1975
- P. andersonii Han et al. 2001
- P. bathycetes Quigley and Colwell 1968
- P. borealis Wilson et al. 2006
- P. cavernae Zhu et al. 2022
- P. cavernicola Zhu et al. 2022
- P. cellulosa Andrews et al. 2000
- P. clemancea Rahman et al. 2010
- P. coenobios ZoBell and Upham 1944
- P. diazotrophicus Watanabe et al. 1987
- P. diterpeniphila Morgan and Wyndham 2002
- P. elodea Fialho et al. 1991
- P. excibis Steinhaus
- P. flexibilis (Hespell 1977) Shin et al. 2016
- P. fluvialis Sudan et al. 2018
- P. gelidicola Kadota 1951 (Approved Lists 1980)
- P. guezennei Simon-Colin et al. 2008
- P. halodenitrificans Alonso et al. 2001
- P. halodurans Cuhel et al. 1981
- P. halosaccharolytica Ohno et al. 1976
- P. halosensibilis Zou and Cai 1994
- P. hydrogenothermophila Goto et al. 1978
- P. hydrogenovora Igarashi et al. 1980
- P. indica Pandey et al. 2002
- P. jinanensis Cai et al. 1989
- P. kuykendallii Hunter and Manter 2012
- P. lopnurensis Mamtimin et al. 2021
- P. lubricans Rehman et al. 2010
- P. matsuisoli Lin et al. 2015
- "P. melophthora Allen and Riker 1932
- P. mesoacidophila Kintaka et al. 1981
- P. multiresinovorans Hernandez et al. 2008
- P. perolens Szybalski 1950
- P. pharmacofabricae corrig. Yu et al. 2019
- P. pratensis Zhang et al. 2021
- P. quercus Li et al. 2021
- P. raguenesii Simon-Colin et al. 2009
- P. reactans Preece and Wong 1982
- P. reptilivora Caldwell and Ryerson 1940
- P. rhizophila Hassen et al. 2018
- P. rhizovicinus He et al. 2021
- P. rubescens Pivnick 1955
- P. schmalbachii Shelomi et al. 2021
- P. septica Bergey et al. 1930
- P. sesami Madhaiyan et al. 2017
- P. siderocapsa Falamin and Pinevich 2006
- P. suis Woods 1930
- P. tamsuii Liang et al. 2015
- P. tarimensis Anwar et al. 2017
- P. teessidea Rahman et al. 2010
- P. thermocarboxydovorans Lyons et al. 1984
- P. thermotolerans Manaia and Moore 2002
- P. tianjinensis Chen et al. 2018
- P. tohonis Yamada et al. 2021
- P. turbinellae Sreenivasan 1956
- P. turukhanskensis Korshunova et al. 2016
- P. tuticorinensis Sreenivasan 1956
- P. wenzhouensis Zhang et al. 2021
- P. xionganensis Zhao et al. 2020
- P. yangonensis Tohya et al. 2020
Species previously classified in the genus[edit]
Recently, 16S rRNA sequence analysis redefined the taxonomy of many bacterial species previously classified as being in the genus Pseudomonas.[11] Species removed from Pseudomonas are listed below; clicking on a species will show its new classification. The term 'pseudomonad' does not apply strictly to just the genus Pseudomonas, and can be used to also include previous members such as the genera Burkholderia and Ralstonia.
α proteobacteria: P. abikonensis, P. aminovorans, P. azotocolligans, P. carboxydohydrogena, P. carboxidovorans, P. compransoris, P. diminuta, P. echinoides, P. extorquens, P. lindneri, P. mesophilica, P. paucimobilis, P. radiora, P. rhodos, P. riboflavina, P. rosea, P. vesicularis.
β proteobacteria: P. acidovorans, P. alliicola, P. antimicrobica, P. avenae, P. butanovora, P. caryophylli, P. cattleyae, P. cepacia, P. cocovenenans, P. delafieldii, P. facilis, P. flava, P. gladioli, P. glathei, P. glumae, P. huttiensis, P. indigofera, P. lanceolata, P. lemoignei, B. mallei, P. mephitica, P. mixta, P. palleronii, P. phenazinium, P. pickettii, P. plantarii, P. pseudoflava, B. pseudomallei, P. pyrrocinia, P. rubrilineans, P. rubrisubalbicans, P. saccharophila, P. solanacearum, P. spinosa, P. syzygii, P. taeniospiralis, P. terrigena, P. testosteroni.
γ-β proteobacteria: P. boreopolis, P. cissicola, P. geniculata, P. hibiscicola, P. maltophilia, P. pictorum.
γ proteobacteria: P. beijerinckii, P. diminuta, P. doudoroffii, P. elongata, P. flectens, P. marinus, P. halophila, P. iners, P. marina, P. nautica, P. nigrifaciens, P. pavonacea,[60] P. piscicida, P. stanieri.
δ proteobacteria: P. formicans.
Phylogenetics[edit]
The following relationships between genomic affinity groups have been determined by phylogenetic analysis:[61][62]
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Bacteriophages[edit]
There are a number of bacteriophages that infect Pseudomonas, e.g.
- Pseudomonas phage Φ6
- Pseudomonas phage ΦCTX
- Pseudomonas aeruginosa phage EL[63]
- Pseudomonas aeruginosa phage ΦKMV (a Phikmvvirus)[64]
- Pseudomonas aeruginosa phage LKD16 (a Phikmvvirus)[65]
- Pseudomonas aeruginosa phage LKA1 (a Phikmvvirus)[65]
- Pseudomonas aeruginosa phage LUZ19 (a Phikmvvirus)
- Pseudomonas aeruginosa phage ΦKZ[63]
- Pseudomonas putida phage gh-1[66]
See also[edit]
- Culture collection for a list of culture collections
Footnotes[edit]
- ^ To aid in the flow of the prose in English, genus names can be "trivialised" to form a vernacular name to refer to a member of the genus: for the genus Pseudomonas it is "pseudomonad" (plural: "pseudomonads"), a variant on the non-nominative cases in the Greek declension of monas, monada.[67] For historical reasons, members of several genera that were formerly classified as Pseudomonas species can be referred to as pseudomonads, while the term "fluorescent pseudomonad" refers strictly to current members of the genus Pseudomonas, as these produce pyoverdin, a fluorescent siderophore.[4][page needed] The latter term, fluorescent pseudomonad, is distinct from the term P. fluorescens group, which is used to distinguish a subset of members of the Pseudomonas sensu stricto and not as a whole
References[edit]
- ^ Lalucat, Jorge; Gomila, Margarita; Mulet, Magdalena; Zaruma, Anderson; García-Valdés, Elena (2021). "Past, present and future of the boundaries of the Pseudomonas genus: Proposal of Stutzerimonas gen. nov". Syst Appl Microbiol. 45 (1): 126289. doi:10.1016/j.syapm.2021.126289. hdl:10261/311157. PMID 34920232. S2CID 244943909.
- ^ Parte, Aidan C.; Sardà Carbasse, Joaquim; Meier-Kolthoff, Jan P.; Reimer, Lorenz C.; Göker, Markus (2020). "List of Prokaryotic names with Standing in Nomenclature (LPSN) moves to the DSMZ". International Journal of Systematic and Evolutionary Microbiology. 70 (11): 5607–5612. doi:10.1099/ijsem.0.004332. PMC 7723251. PMID 32701423.
- ^ "Genus Pseudomonas". LPSN.dsmz.de. Retrieved 4 April 2023. Partial citation, see Parte et al., 2020 for project reference
- ^ a b Madigan M; Martinko J, eds. (2006). Brock Biology of Microorganisms (11th ed.). Prentice Hall. ISBN 0-13-144329-1.
- ^ Padda, Kiran Preet; Puri, Akshit; Chanway, Chris (2019-11-01). "Endophytic nitrogen fixation – a possible 'hidden' source of nitrogen for lodgepole pine trees growing at unreclaimed gravel mining sites". FEMS Microbiology Ecology. 95 (11). doi:10.1093/femsec/fiz172. ISSN 0168-6496. PMID 31647534.
- ^ Padda, Kiran Preet; Puri, Akshit; Chanway, Chris P. (2018-09-20). "Isolation and identification of endophytic diazotrophs from lodgepole pine trees growing at unreclaimed gravel mining pits in central interior British Columbia, Canada". Canadian Journal of Forest Research. 48 (12): 1601–1606. doi:10.1139/cjfr-2018-0347. hdl:1807/92505. ISSN 0045-5067. S2CID 92275030.
- ^ Migula, W. (1894) Über ein neues System der Bakterien. Arb Bakteriol Inst Karlsruhe 1: 235–238.
- ^ Migula, W. (1900) System der Bakterien, Vol. 2. Jena, Germany: Gustav Fischer.
- ^ a b c Palleroni, N. J. (2010). "The Pseudomonas Story". Environmental Microbiology. 12 (6): 1377–1383. Bibcode:2010EnvMi..12.1377P. doi:10.1111/j.1462-2920.2009.02041.x. PMID 20553550.
- ^ a b c Cornelis, Pierre, ed. (2008). Pseudomonas: Genomics and Molecular Biology (1st ed.). Caister Academic Press. ISBN 978-1-904455-19-6.
- ^ a b Anzai, Y.; Kim, H.; Park, J. Y.; Wakabayashi, H. (2000). "Phylogenetic affiliation of the pseudomonads based on 16S rRNA sequence". International Journal of Systematic and Evolutionary Microbiology. 50 (4): 1563–89. doi:10.1099/00207713-50-4-1563. PMID 10939664.
- ^ Anzai, Yojiro; Kudo, Yuko; Oyaizu, Hiroshi (1997). "The phylogeny of the genera Chryseomonas, Flavimonas, and Pseudomonas supports synonymy of these three genera". International Journal of Systematic Bacteriology. 47 (2): 249–251. doi:10.1099/00207713-47-2-249. PMID 9103607.
- ^ Yabuuchi, Eiko; Kosako, Yoshimasa; Oyaizu, Hiroshi; Yano, Ikuya; Hotta, Hisako; Hashimoto, Yasuhiro; Ezaki, Takayuki; Arakawa, Michio (1992). "Proposal of Burkholderia gen. Nov. And transfer of seven species of the genus Pseudomonas homology group II to the new genus, with the type species Burkholderia cepacia (Palleroni and Holmes 1981) comb. Nov". Microbiology and Immunology. 36 (12): 1251–1275. doi:10.1111/j.1348-0421.1992.tb02129.x. PMID 1283774. S2CID 46648461.
- ^ Yabuuchi, Eiko; Kosako, Yoshimasa; Yano, Ikuya; Hotta, Hisako; Nishiuchi, Yukiko (1995). "Transfer of two Burkholderia and an Alcaligenes species to Ralstonia gen. Nov.: Proposal of Ralstonia pickettii (Ralston, Palleroni and Doudoroff 1973) comb. Nov., Ralstonia solanacearum (Smith 1896) comb. Nov. And Ralstonia eutropha (Davis 1969) comb. Nov". Microbiology and Immunology. 39 (11): 897–904. doi:10.1111/j.1348-0421.1995.tb03275.x. PMID 8657018. S2CID 28187828.
- ^ a b c d e f Nikolaidis, Marios; Mossialos, Dimitris; Oliver, Stephen G.; Amoutzias, Grigorios D. (2020-07-24). "Comparative Analysis of the Core Proteomes among the Pseudomonas Major Evolutionary Groups Reveals Species-Specific Adaptations for Pseudomonas aeruginosa and Pseudomonas chlororaphis". Diversity. 12 (8): 289. doi:10.3390/d12080289. ISSN 1424-2818.
- ^ Richter, Michael; Rosselló-Móra, Ramon (2009-11-10). "Shifting the genomic gold standard for the prokaryotic species definition". Proceedings of the National Academy of Sciences. 106 (45): 19126–19131. Bibcode:2009PNAS..10619126R. doi:10.1073/pnas.0906412106. ISSN 0027-8424. PMC 2776425. PMID 19855009.
- ^ Tran, Phuong N.; Savka, Michael A.; Gan, Han Ming (2017-07-12). "In-silico Taxonomic Classification of 373 Genomes Reveals Species Misidentification and New Genospecies within the Genus Pseudomonas". Frontiers in Microbiology. 8: 1296. doi:10.3389/fmicb.2017.01296. ISSN 1664-302X. PMC 5506229. PMID 28747902.
- ^ a b Koehorst, Jasper J.; Dam, Jesse C. J.; van Heck, Ruben G. A.; van Saccenti, Edoardo; Martins dos Santos, Vitor A. P.; Suarez-Diez, Maria; Schaap, Peter J. (2016-12-06). "Comparison of 432 Pseudomonas strains through integration of genomic, functional, metabolic and expression data". Scientific Reports. 6 (1): 38699. Bibcode:2016NatSR...638699K. doi:10.1038/srep38699. ISSN 2045-2322. PMC 5138606. PMID 27922098.
- ^ Krieg, Noel (1984). Bergey's Manual of Systematic Bacteriology, Volume 1. Baltimore: Williams & Wilkins. ISBN 0-683-04108-8.
- ^ Meyer, Jean-Marie; Geoffroy, Valérie A.; Baida, Nader; Gardan, Louis; Izard, Daniel; Lemanceau, Philippe; Achouak, Wafa; Palleroni, Norberto J. (2002). "Siderophore typing, a powerful tool for the identification of fluorescent and nonfluorescent pseudomonads". Applied and Environmental Microbiology. 68 (6): 2745–2753. Bibcode:2002ApEnM..68.2745M. doi:10.1128/AEM.68.6.2745-2753.2002. PMC 123936. PMID 12039729.
- ^ Lau, Gee W.; Hassett, Daniel J.; Ran, Huimin; Kong F, Fansheng (2004). "The role of pyocyanin in Pseudomonas aeruginosa infection". Trends in Molecular Medicine. 10 (12): 599–606. doi:10.1016/j.molmed.2004.10.002. PMID 15567330.
- ^ Matthijs, Sandra; Tehrani, Kourosch Abbaspour; Laus, George; Jackson, Robert W.; Cooper, Richard M.; Cornelis, Pierre (2007). "Thioquinolobactin, a Pseudomonas siderophore with antifungal and anti-Pythium activity". Environmental Microbiology. 9 (2): 425–434. Bibcode:2007EnvMi...9..425M. doi:10.1111/j.1462-2920.2006.01154.x. PMID 17222140.
- ^ Biello, David (February 28, 2008). "Do Microbes Make Snow?". Scientific American.
- ^ Hassett, Daniel J.; Cuppoletti, John; Trapnell, Bruce; Lymar, Sergei V.; et al. (2002). "Anaerobic metabolism and quorum sensing by Pseudomonas aeruginosa biofilms in chronically infected cystic fibrosis airways: rethinking antibiotic treatment strategies and drug targets". Advanced Drug Delivery Reviews. 54 (11): 1425–1443. doi:10.1016/S0169-409X(02)00152-7. PMID 12458153.
- ^ a b Ryan, Kenneth J.; Ray, C. George; Sherris, John C., eds. (2004). Sherris Medical Microbiology (4th ed.). McGraw Hill. ISBN 0-8385-8529-9.
- ^ Van Eldere, Johan (February 2003). "Multicentre surveillance of Pseudomonas aeruginosa susceptibility patterns in nosocomial infections". Journal of Antimicrobial Chemotherapy. 51 (2): 347–352. doi:10.1093/jac/dkg102. PMID 12562701.
- ^ Poole, K (January 2004). "Efflux-mediated multiresistance in Gram-negative bacteria". Clinical Microbiology and Infection. 10 (1): 12–26. doi:10.1111/j.1469-0691.2004.00763.x. PMID 14706082.
- ^ "Scientists Discover Clays to Fight Deadly Bacteria". www.infoniac.com. 2007-03-16. Retrieved 2008-11-20.
- ^ Smith, Michael (2007-03-16). "Gallium May Have Antibiotic-Like Properties". MedPage Today. Archived from the original on 2008-09-18.
- ^ Bodey, Gerald P.; Bolivar, Ricardo; Fainstein, Victor; Jadeja, Leena (1983-03-01). "Infections Caused by Pseudomonas aeruginosa". Clinical Infectious Diseases. 5 (2): 279–313. doi:10.1093/clinids/5.2.279. ISSN 1058-4838. PMID 6405475.
- ^ Hardie, Kim R.; Pommier, Stephanie; Wilhelm, Susanne (2009). "The Secreted Proteins of Pseudomonas aeruginosa: Their Export Machineries, and How They Contribute to Pathogenesis". Bacterial Secreted Proteins: Secretory Mechanisms and Role in Pathogenesis. Caister Academic Press. ISBN 978-1-904455-42-4.
- ^ Brodey, Catherine L.; Rainey, Paul B.; Tester, Mark; Johnstone, Keith (1991). "Bacterial blotch disease of the cultivated mushroom is caused by an ion channel forming lipodepsipeptide toxin". Molecular Plant-Microbe Interactions. 1 (4): 407–411. doi:10.1094/MPMI-4-407.
- ^ Young, J. M. (1970). "Drippy gill: a bacterial disease of cultivated mushrooms caused by Pseudomonas agarici n. sp". New Zealand Journal of Agricultural Research. 13 (4): 977–90. Bibcode:1970NZJAR..13..977Y. doi:10.1080/00288233.1970.10430530.
- ^ Haas, Dieter; Défago, Geneviève (2005). "Biological control of soil-borne pathogens by fluorescent pseudomonads". Nature Reviews Microbiology. 3 (4): 307–319. doi:10.1038/nrmicro1129. PMID 15759041. S2CID 18469703.
- ^ Chin-A-Woeng TF; Bloemberg, Guido V.; Mulders, Ine H. M.; Dekkers, Linda C.; et al. (2000). "Root colonization by phenazine-1-carboxamide-producing bacterium Pseudomonas chlororaphis PCL1391 is essential for biocontrol of tomato foot and root rot". Mol Plant Microbe Interact. 13 (12): 1340–1345. doi:10.1094/MPMI.2000.13.12.1340. hdl:1887/62881. PMID 11106026.
- ^ Esipov, SE; Adanin, VM; Baskunov, BP; Kiprianova, EA; et al. (1975). "Novyĭ antibioticheski aktivnyĭ florogliutsid iz Pseudomonas aurantiaca" [New antibiotically active fluoroglucide from Pseudomonas aurantiaca]. Antibiotiki (in Russian). 20 (12): 1077–81. PMID 1225181.
- ^ O’Mahony, Mark M.; Dobson, Alan D. W.; Barnes, Jeremy D.; Singleton, Ian (2006). "The use of ozone in the remediation of polycyclic aromatic hydrocarbon contaminated soil". Chemosphere. 63 (2): 307–314. Bibcode:2006Chmsp..63..307O. doi:10.1016/j.chemosphere.2005.07.018. PMID 16153687.
- ^ Yen, K M; Karl, M R; Blatt, L M; Simon, M J; Winter, R B; Fausset, P R; Lu, H S; Harcourt, A A; Chen, K K (1991). "Cloning and characterization of a Pseudomonas mendocina KR1 gene cluster encoding toluene-4-monooxygenase". Journal of Bacteriology. 173 (17): 5315–27. doi:10.1128/jb.173.17.5315-5327.1991. PMC 208241. PMID 1885512.
- ^ Huertas, M.-J.; Luque-Almagro, V.M.; Martínez-Luque, M.; Blasco, R.; Moreno-Vivián, C.; Castillo, F.; Roldán, M.-D. (2006). "Cyanide metabolism of Pseudomonas pseudoalcaligenes CECT5344: role of siderophores". Biochemical Society Transactions. 34 (1): 152–5. doi:10.1042/BST0340152. PMID 16417508.
- ^ Nojiri, Hideaki; Maeda, Kana; Sekiguchi, Hiroyo; Urata, Masaaki; Shintani, Masaki; Yoshida, Takako; Habe, Hiroshi; Omori, Toshio (2002). "Organization and transcriptional characterization of catechol degradation genes involved in carbazole degradation by Pseudomonas resinovorans strain CA10". Bioscience, Biotechnology, and Biochemistry. 66 (4): 897–901. doi:10.1271/bbb.66.897. PMID 12036072.
- ^ Gilani, Razia Alam; Rafique, Mazhar; Rehman, Abdul; Munis, Muhammad Farooq Hussain; Rehman, Shafiq ur; Chaudhary, Hassan Javed (2016). "Biodegradation of chlorpyrifos by bacterial genus Pseudomonas". Journal of Basic Microbiology. 56 (2): 105–119. doi:10.1002/jobm.201500336. ISSN 1521-4028. PMID 26837064. S2CID 1373984.
- ^ Nam, IH; Chang, YS; Hong, HB; Lee, YE (2003). "A novel catabolic activity of Pseudomonas veronii in biotransformation of pentachlorophenol". Applied Microbiology and Biotechnology. 62 (2–3): 284–290. doi:10.1007/s00253-003-1255-1. PMID 12883877. S2CID 31700132.
- ^ Onaca, Christina; Kieninger, Martin; Engesser, Karl H.; Altenbuchner, Josef (May 2007). "Degradation of alkyl methyl ketones by Pseudomonas veronii". Journal of Bacteriology. 189 (10): 3759–3767. doi:10.1128/JB.01279-06. PMC 1913341. PMID 17351032.
- ^ Marqués, Silvia; Ramos, Juan L. (1993). "Transcriptional control of the Pseudomonas putida TOL plasmid catabolic pathways". Molecular Microbiology. 9 (5): 923–929. doi:10.1111/j.1365-2958.1993.tb01222.x. PMID 7934920. S2CID 20663917.
- ^ Sepúlveda-Torres, Lycely Del C.; Rajendran, Narayanan; Dybas, Michael J.; Criddle, Craig S. (1999). "Generation and initial characterization of Pseudomonas stutzeri KC mutants with impaired ability to degrade carbon tetrachloride". Archives of Microbiology. 171 (6): 424–429. Bibcode:1999ArMic.171..424D. doi:10.1007/s002030050729. PMID 10369898. S2CID 19916486.
- ^ "What Is Pseudomonas Aeruginosa?". WebMD. October 27, 2022. Retrieved 2023-08-07.
- ^ a b c Wood, Peter (2021-03-16). "Pseudomonas: How to Treat and Prevent in Commercial Water Systems". Wychwood Water Systems. Retrieved 2023-08-07.
- ^ a b c d Dasen, S. E.; LiPuma, J. J.; Kostman, J. R.; Stull, T. L. (1 October 1994). "Characterization of PCR-ribotyping for Burkholderia (Pseudomonas) cepacia". Journal of Clinical Microbiology. 32 (10): 2422–2424. doi:10.1128/JCM.32.10.2422-2424.1994. ISSN 0095-1137. PMC 264078. PMID 7529239.
- ^ a b c d e Dogan, Belgin; Boor, Kathryn J. (1 January 2003). "Genetic Diversity and Spoilage Potentials among Pseudomonas spp. Isolated from Fluid Milk Products and Dairy Processing Plants". Applied and Environmental Microbiology. 69 (1): 130–138. Bibcode:2003ApEnM..69..130D. doi:10.1128/AEM.69.1.130-138.2003. ISSN 0099-2240. PMC 152439. PMID 12513987.
- ^ a b c Casalinuovo, Ida A.; Di Pierro, Donato; Coletta, Massimiliano; Di Francesco, Paolo (1 November 2006). "Application of Electronic Noses for Disease Diagnosis and Food Spoilage Detection". Sensors. 6 (11): 1428–1439. Bibcode:2006Senso...6.1428C. doi:10.3390/s6111428. PMC 3909407.
- ^ a b c d Magan, Naresh; Pavlou, Alex; Chrysanthakis, Ioannis (5 January 2001). "Milk-sense: a volatile sensing system recognises spoilage bacteria and yeasts in milk". Sensors and Actuators B: Chemical. 72 (1): 28–34. doi:10.1016/S0925-4005(00)00621-3.
- ^ Pseudomonas in LPSN; Parte, Aidan C.; Sardà Carbasse, Joaquim; Meier-Kolthoff, Jan P.; Reimer, Lorenz C.; Göker, Markus (1 November 2020). "List of Prokaryotic names with Standing in Nomenclature (LPSN) moves to the DSMZ". International Journal of Systematic and Evolutionary Microbiology. 70 (11): 5607–5612. doi:10.1099/ijsem.0.004332.
- ^ Anzai, Y; Kim, H; Park, J Y; Wakabayashi, H; Oyaizu, H (2000). "Phylogenetic affiliation of the pseudomonads based on 16S rRNA sequence". International Journal of Systematic and Evolutionary Microbiology. 50 (4): 1563–1589. doi:10.1099/00207713-50-4-1563. ISSN 1466-5034. PMID 10939664.
- ^ Jun, Se-Ran; Wassenaar, Trudy M.; Nookaew, Intawat; Hauser, Loren; Wanchai, Visanu; Land, Miriam; Timm, Collin M.; Lu, Tse-Yuan S.; Schadt, Christopher W.; Doktycz, Mitchel J.; Pelletier, Dale A.; Ussery, David W. (2016). "Diversity of Pseudomonas Genomes, Including Populus-Associated Isolates, as Revealed by Comparative Genome Analysis". Applied and Environmental Microbiology. 82 (1): 375–383. Bibcode:2016ApEnM..82..375J. doi:10.1128/AEM.02612-15. ISSN 0099-2240. PMC 4702629. PMID 26519390.
- ^ Mulet, Magdalena; Lalucat, Jorge; García-Valdés, Elena (2010). "DNA sequence-based analysis of the Pseudomonas species". Environmental Microbiology. 12 (6): 1513–30. Bibcode:2010EnvMi..12.1513M. doi:10.1111/j.1462-2920.2010.02181.x. PMID 20192968.
- ^ Mulet, Magdalena; Gomila, Margarita; Scotta, Claudia; Sánchez, David; Lalucat, Jorge; García-Valdés, Elena (2012). "Concordance between whole-cell matrix-assisted laser-desorption/ionization time-of-flight mass spectrometry and multilocus sequence analysis approaches in species discrimination within the genus Pseudomonas". Systematic and Applied Microbiology. 35 (7): 455–464. doi:10.1016/j.syapm.2012.08.007. ISSN 0723-2020. PMID 23140936.
- ^ Gomila, Margarita; Peña, Arantxa; Mulet, Magdalena; Lalucat, Jorge; García-Valdés, Elena (2015). "Phylogenomics and systematics in Pseudomonas". Frontiers in Microbiology. 6: 214. doi:10.3389/fmicb.2015.00214. ISSN 1664-302X. PMC 4447124. PMID 26074881.
- ^ Hesse, Cedar; Schulz, Frederik; Bull, Carolee T.; Shaffer, Brenda T.; Yan, Qing; Shapiro, Nicole; Hassan, Karl A.; Varghese, Neha; Elbourne, Liam D. H.; Paulsen, Ian T.; Kyrpides, Nikos; Woyke, Tanja; Loper, Joyce E. (2018). "Genome-based evolutionary history of Pseudomonas spp". Environmental Microbiology. 20 (6): 2142–2159. Bibcode:2018EnvMi..20.2142H. doi:10.1111/1462-2920.14130. ISSN 1462-2912. OSTI 1529110. PMID 29633519. S2CID 4737911.
- ^ Girard, Léa; Lood, Cédric; Höfte, Monica; Vandamme, Peter; Rokni-Zadeh, Hassan; van Noort, Vera; Lavigne, Rob; De Mot, René (2021). "The Ever-Expanding Pseudomonas Genus: Description of 43 New Species and Partition of the Pseudomonas putida Group". Microorganisms. 9 (8): 1766. doi:10.3390/microorganisms9081766. ISSN 2076-2607. PMC 8401041. PMID 34442845.
- ^ Van Landschoot, A.; Rossau, R.; De Ley, J. (1986). "Intra- and Intergeneric Similarities of the Ribosomal Ribonucleic Acid Cistrons of Acinetobacter". International Journal of Systematic Bacteriology. 36 (2): 150. doi:10.1099/00207713-36-2-150.
- ^ Girard L, Lood C, Höfte M, Vandamme P, Rokni-Zadeh H, van Noort V, Lavigne R, De Mot R. (2021). "The Ever-Expanding Pseudomonas Genus: Description of 43 New Species and Partition of the Pseudomonas putida Group". Microorganisms. 9 (8): 1766. doi:10.3390/microorganisms9081766. PMC 8401041. PMID 34442845.
- ^ Yi B, Dalpke AH. (2022). "Revisiting the intrageneric structure of the genus Pseudomonas with complete whole genome sequence information: Insights into diversity and pathogen-related genetic determinants". Infect Genet Evol. 97: 105183. doi:10.1016/j.meegid.2021.105183. PMID 34920102. S2CID 245180021. Note that the tree in this reference has the same topology, but looks different because it is unrooted.
- ^ a b Hertveldt, K.; Lavigne, R.; Pleteneva, E.; Sernova, N.; Kurochkina, L.; Korchevskii, R.; Robben, J.; Mesyanzhinov, V.; Krylov, V. N.; Volckaert, G. (2005). "Genome Comparison of Pseudomonas aeruginosa Large Phages" (PDF). Journal of Molecular Biology. 354 (3): 536–545. doi:10.1016/j.jmb.2005.08.075. PMID 16256135. Archived from the original (PDF) on 2016-03-04. Retrieved 2015-08-27.
- ^ Lavigne, Rob; Noben, Jean-Paul; Hertveldt, Kirsten; Ceyssens, Pieter-Jan; Briers, Yves; Dumont, Debora; Roucourt, Bart; Krylov, Victor N.; Mesyanzhinov, Vadim V.; Robben, Johan; Volckaert, Guido (2006). "The structural proteome of Pseudomonas aeruginosa bacteriophage ϕKMV". Microbiology. 152 (2): 529–534. doi:10.1099/mic.0.28431-0. ISSN 1465-2080. PMID 16436440.
- ^ a b Ceyssens, Pieter-Jan; Lavigne, Rob; Mattheus, Wesley; Chibeu, Andrew; Hertveldt, Kirsten; Mast, Jan; Robben, Johan; Volckaert, Guido (2006). "Genomic Analysis of Pseudomonas aeruginosa Phages LKD16 and LKA1: Establishment of the φKMV Subgroup within the T7 Supergroup". Journal of Bacteriology. 188 (19): 6924–6931. doi:10.1128/JB.00831-06. ISSN 0021-9193. PMC 1595506. PMID 16980495.
- ^ Lee, Lucy F.; Boezi, J. A. (1966). "Characterization of bacteriophage gh-1 for Pseudomonas putida". Journal of Bacteriology. 92 (6). American Society for Microbiology: 1821–1827. doi:10.1128/JB.92.6.1821-1827.1966. PMC 316266. PMID 5958111.
- ^ Buchanan, R. E. (1955). "Taxonomy". Annual Review of Microbiology. 9: 1–20. doi:10.1146/annurev.mi.09.100155.000245. PMID 13259458.
