Resolvin
This article needs more reliable medical references for verification or relies too heavily on primary sources. (November 2017) |  |
This article may be too technical for most readers to understand. (March 2018) |
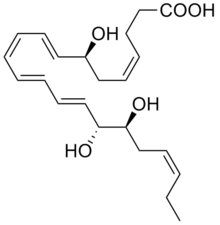
Resolvins are specialized pro-resolving mediators (SPMs) derived from omega-3 fatty acids, primarily eicosapentaenoic acid (EPA) and docosahexaenoic acid (DHA), as well as from two isomers of docosapentaenoic acid (DPA), one omega-3 and one omega-6 fatty acid. As autacoids similar to hormones acting on local tissues, resolvins are under preliminary research for their involvement in promoting restoration of normal cellular function following the inflammation that occurs after tissue injury.[1][2] Resolvins belong to a class of polyunsaturated fatty acid (PUFA) metabolites termed specialized proresolving mediators (SPMs).[3][4]
Biochemistry and production[edit]
Resolvins (Rvs) fall into several sub-classes based on the straight chain PUFA from which they are formed and derive their unique structure. The resolvins Ds (RvDs) are metabolites of the 22-carbon PUFA, DHA (i.e. 4Z,7Z,10Z,13Z,16Z,19Z-docosahexaenoic acid); the resolvins Es (RvEs) are metabolites of the 20-carbon PUFA, EPA (i.e. 5Z,8Z,11Z,14Z,17Z-eicosapentaenoic acid); the resolvins Dn-6DPA (RvDsn-6DPA) are metabolites of the DPA isomer, osbond acid (i.e. 4Z,7Z,10Z,13Z,16Z-docosapentaenoic acid); the resolvins Dn-3DPA (RvDn-3DPA) are metabolites of the DPA isomer, clupanodonic acid (i.e. 7Z,10Z,13Z,16Z,19Z-docosapentaenoic acid); and the resolvins Ts (RvTs) are metabolites of clupanodonic acid, that possess a 17R hydroxyl residue, whereas all RvDsn-3DPA resolvins have a 17S hydroxyl residue. Certain isomers of RvDs are termed aspirin-triggered resolvin Ds (AT-RvDs) because their synthesis is initiated by a drug-modified COX2 enzyme to form 17(R) hydroxyl rather than 17(S) hydroxyl residue of the RvEs; however, an unidentified as of 2023 cytochrome P450 enzyme(s) may also form this 17(R)-hydroxy intermediate and thereby contribute to the production of AT-RvEs. All of the cited resolvins except the RvDsn-6DPA are metabolites of omega-3 fatty acids.[3][4]
The following oxygenase enzymes may be responsible for metabolizing PUFA to resolvins: 15-lipoxygenase-1 (i.e. ALOX15), possibly 15-lipoxygenase-2 (i.e. ALOX15B), 5-lipoxygenase (i.e. ALOX5), cyclooxygenase-2 (i.e. COX-2), and certain cytochrome P450 monooxygenases.[3][5]
Resolvin Ds[edit]
RvDs are poly-hydroxyl metabolites of DHA. To date, six RvDs, which vary in the number, position, and chirality of their hydroxyl residues as well as the position and cis–trans isomerism of their 6 double bonds, have been described. These are: RvD1 (7S,8R,17S-trihydroxy-DHA), RvD2 (7S,16R,17S-trihydroxy-DHA), RvD3 (4S,7R,17S-trihydroxy-DHA), RvD4 (4S,5,17S-trihydroxy-DHA; chirality at position 5 not yet determined as of 2023), RvD5 (7S,17S-dihydroxy-DHA), and RvD6 (4S,17S-dihydroxy-DHA). (The structures of these RvDs are further defined at Specialized pro-resolving mediators § DHA-derived resolvins). These metabolites are formed by a wide range of cells and tissues by the initial metabolism of DHA to 7S-hydroperoxy-DHA and 4S-hydroperoxy-DHA by a 15-lipoxygenase (either ALOX15 or possibly ALOX15B) followed by the further metabolism of the two intermediates by ALOX5 to their 17-hydroperoxy derivatives; these di-hydroperoxy products are further altered to the cited RvDs by these oxygenases or by non-enzymatic reactions and the conversion of their peroxy residues ubiquitous cellular peroxidases.[3][5]
Resolvin Es[edit]
RvEs are di- or tri-hydroxyl metabolites of EPA. To date, four RvEs have been described: RvE1 (5S,12R,18R-trihydroxy-EPA), 18S-Rv1 (5S,12R,18S-trihydroxy-EPA), RvE2 (5S,18R-dihydroxy-EPA), and RvE3 (17R,18R/S-dihydroxy-EPA). (Structures of the RvEs are further defined at Specialized pro-resolving mediators § EPA-derived resolvins). Resolvins Es are formed in manner similar to AT resolvins Ts. COX-2 modified in activity by aspirin or atorvastatin or, alternatively, a microbial or possibly mammalian cytochrome P450 monoxygenase metabolizes EPA to its 18R-hydroperoxy derivative; this intermediate is then further metabolized by ALOX5 to a 5,6 epoxide which is hydrolyzed enzymatically or non-enzymatically to RvE1 and 18S-RvE1 or reduced to RvE2; alternatively the 18R-hydroperoxide is converted to the 17R,18S vicinal diol product, RvE3.[3][5]
T series resolvins[edit]
Human platelets pretreated with aspirin or atorvastatin metabolize the omega-3 DPA, clupanodonic acid (DPAn-3) by aspirin-treated or atorvastatin-treated COX-2 to a 13S-hydroperoxy intermediate (aspirin and atorvastatin change the activity of COX-2 from a cyclooxygenase to a hydroxyperoxidase-forming enzyme. The intermediate is then passed to nearby human neutrophils which metabolize it, probably by ALOX5 enzyme activity, to four poly-hydroxyl metabolites: RvT1 (7S,13R,20S-trihydroxy-8E,10Z,14E,16Z,18E-DPA), RvT2 (7S,12R,13S-trihydroxy-8Z,10E,14E,16Z,19Z-DPA), RvT3 (7S,8R,13S-trihydroxy-9E,11E,14E,16Z,19Z-DPA), and RvT4 (7S,13R-dihydroxy-8E,10Z,14E,16Z,19Z-DPA).[6] Subsequent studies found that these four RvTs are also formed by mixtures of human neutrophils and vascular endothelium cells and, additionally, are detected in the infected tissues of rodents and humans.[7][8]
Putative mechanisms[edit]
This section needs more reliable medical references for verification or relies too heavily on primary sources. (March 2018) |  |
Following tissue injury, the inflammatory response is a protective process to promote restoration of the tissue to homeostasis.[2] Resolution of inflammation involves various specialized lipid mediators, including resolvins.[1][2] Resolvins are under laboratory research for their potential to act through G protein-coupled receptors (GPRs): 1) RvD1 and AT-RvD1 act through the formyl peptide receptor 2, which is also activated by certain lipoxins and is therefore often termed the ALX/FPR2 receptor; 2) RvD1, AT-RVD1, RvD3, AT-RvD3, and RvD5 act through the GPR32 receptor which is now also termed the RVD1 receptor; 3) RvD2 acts through the GPR18 receptor also now termed the RvD2 receptor; and 4) RvE1 and the 18(S) analog of RvE1 are full activators while RvE2 is a partial activator of the CMKLR1 receptor. All of these receptors activate their parent cells through standard GPR-mobilized pathways.[4][9] RvE1, 18(S)-RvE1, and RvE2 inhibit the leukotriene B4 receptor 1 which is the receptor for inflammation-promoting PUFA metabolites such as LTB4 and the R stereoisomer of 12-HETE; by inhibiting the action of these pro-inflammatory mediators.[5][9]
References[edit]
- ^ a b Moro, K; Nagahashi, M; Ramanathan, R; Takabe, K; Wakai, T (2016). "Resolvins and omega three polyunsaturated fatty acids: Clinical implications in inflammatory diseases and cancer". World Journal of Clinical Cases. 4 (7): 155–164. doi:10.12998/wjcc.v4.i7.155. PMC 4945585. PMID 27458590.
- ^ a b c Balta, M. G; Loos, B. G; Nicu, E. A (2017). "Emerging Concepts in the Resolution of Periodontal Inflammation: A Role for Resolvin E1". Frontiers in Immunology. 8: 1682. doi:10.3389/fimmu.2017.01682. PMC 5735081. PMID 29312286.
- ^ a b c d e Serhan, C. N.; Chiang, N; Dalli, J; Levy, B. D. (2014). "Lipid mediators in the resolution of inflammation". Cold Spring Harbor Perspectives in Biology. 7 (2): a016311. doi:10.1101/cshperspect.a016311. PMC 4315926. PMID 25359497.
- ^ a b c Duvall, M. G.; Levy, B. D. (2015). "DHA- and EPA-derived resolvins, protectins, and maresins in airway inflammation". European Journal of Pharmacology. 785: 144–155. doi:10.1016/j.ejphar.2015.11.001. PMC 4854800. PMID 26546247.
- ^ a b c d Qu Q, Xuan W, Fan GH (2015). "Roles of resolvins in the resolution of acute inflammation". Cell Biology International. 39 (1): 3–22. doi:10.1002/cbin.10345. PMID 25052386. S2CID 10160642.
- ^ Liu C, Fan D, Lei Q, Lu A, He X (December 2022). "Roles of Resolvins in Chronic Inflammatory Response". International Journal of Molecular Sciences. 23 (23). Table 1. doi:10.3390/ijms232314883. PMC 9738788. PMID 36499209.
- ^ Dalli J, Colas RA, Serhan CN (2013). "Novel n-3 immunoresolvents: structures and actions". Scientific Reports. 3: 1940. Bibcode:2013NatSR...3E1940D. doi:10.1038/srep01940. PMC 3672887. PMID 23736886.
- ^ Dalli J, Chiang N, Serhan CN (2015). "Elucidation of novel 13-series resolvins that increase with atorvastatin and clear infections" (PDF). Nature Medicine. 21 (9): 1071–1075. doi:10.1038/nm.3911. PMC 4560998. PMID 26236990.
- ^ a b Serhan, C. N. (2014). "Pro-resolving lipid mediators are leads for resolution physiology". Nature. 510 (7503): 92–101. Bibcode:2014Natur.510...92S. doi:10.1038/nature13479. PMC 4263681. PMID 24899309.
