TMEM249
| TMEM249 | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | TMEM249, C8ORFK29, TMEM 249, transmembrane protein 249 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | MGI: 3647471; HomoloGene: 90450; GeneCards: TMEM249; OMA:TMEM249 - orthologs | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
TMEM249 is a protein that in humans is encoded by the C8orfk29 gene.
Locus[edit]
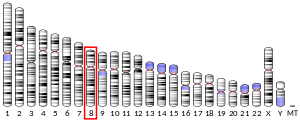
TMEM249 is located near the end of the long arm of chromosome 8 in humans.[5]

Common aliases[edit]
TMEM249 is also known as C8orfk29.[6]
Primary sequence and variants or isoforms[edit]

The primary sequence found at NCBI[7] and Aceview on NCBI predicts there are five spliceforms, with four closely resembling one another and the fifth missing a large 5' intron region. Softberry reinforces the Aceview data by predicting five exons, which are seen in four of the five spliceforms of Aceview.
The general structure of the TMEM249 transcript has a large 5' UTR followed by exon 1, then a large intron followed by exon 2, a small intron then exon 3. The rest of the protein follows exon 3 with a large intron, exon 4 a small intron then exon 5, the 3' UTR.
The primary transcript contains all five exons and produces a protein that is 235 amino acids long. Transcript 1 and 2 are translated in the 3' to 5' direction while transcripts 3 through 5 are translated in the 5' to 3' direction. The gene is encoded on the minus strand within the chromosome.
Paralogs[edit]

The only known paralog of human TMEM249 is found in the second isoform of the protein in gorillas. Of the 217 amino acids aligned between gorilla and human TMEM249, 96% are in complete consensus and 99% are conserved.
Orthologs[edit]
TMEM249 orthologs includes all groups of life except birds, fungi, archea, protists, and plants. The most distant ortholog, Rozella allomycis, was the most diverged species that qualified as an ortholog. The last shared ancestor between Rozella allomycis and Homo sapiens is the Opisthokonts.
Homologs[edit]
No homologs or homologous domains exist within TMEM249.
Phylogeny[edit]
No fungi orthologs were found in the search for similar sequences, so it could be assumed that the gene may have arisen in Opisthokonts and proliferated down the animal tree. This would mean the protein diverged too late to evolve through the fungi tree. This would explain why there are no found plant orthologs as the gene would have arisen after animals and plants diverged evolutionarily.[citation needed]
Domains and motifs[edit]

There are three predicted transmembrane domains. It is unknown whether these transmembrane domains affect the larger structure of the protein complex once properly expressed in tissue. Evolutionary analysis showed that these transmembrane domains are highly conserved across all ortholog taxa.
Post-translational modifications[edit]

There is an area near the 3’ end of the protein that is predicted to be heavily serine phosphorylated. This end of the protein is likely on the cytosolic half of the protein and serves in some activation function of a pathway.
Secondary structure[edit]

TMEM249 has a highly varied structure. Prediction data supports alternating regions of beta sheets and alpha helices. These predictions may support a beta barrel or "helix barrel" through the membrane made up of multiple protein monomers of TMEM249.
Promoter[edit]

The promoter region was found using Eldorado from Genomatix.de (source), the region occupies a region upstream of the 5’ region of TMEM249 on the minus strand of chromosome 8. This promoter binds a number of transcription factors as determined by Eldorado at Genomatix.de.
Tissue Expression[edit]

TMEM249 expression is present at a high level in a wide variety of human tissues. GEO tissue profiles for this protein show that this protein is present in a wide variety of locations within the human body. The human protein atlas claims an even wider expression scope for this protein(source).
Interacting Proteins[edit]
There were no known protein interactions for TMEM249.
Clinical Significance[edit]
TMEM249 has no known link to medical disease.
Mutations[edit]

There exist a number of SNPs for TMEM249 in humans. The mutations are scattered for the most part, with the largest changes in amino acids occurring in the domain of unknown function.
References[edit]
- ^ a b c ENSG00000285272 GRCh38: Ensembl release 89: ENSG00000261587, ENSG00000285272 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000116376 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "TMEM249 transmembrane protein 249 [Homo sapiens (human)] - Gene - NCBI".
- ^ "TMEM249 Gene - GeneCards | TM249 Protein | TM249 Antibody".
- ^ "Homo sapiens chromosome 8, GRCh38.p13 Primary Assembly". 2019-06-14.
{{cite journal}}: Cite journal requires|journal=(help)
Further reading[edit]
- Strausberg RL, Feingold EA, Grouse LH, Derge JG, Klausner RD, Collins FS, et al. (December 2002). "Generation and initial analysis of more than 15,000 full-length human and mouse cDNA sequences". Proceedings of the National Academy of Sciences of the United States of America. 99 (26): 16899–903. Bibcode:2002PNAS...9916899M. doi:10.1073/pnas.242603899. PMC 139241. PMID 12477932.