User:Eve1359/sandbox
Distribution of fitness effects[edit]
In reality, viewing the fitness effects of mutations in these discrete categories is an oversimplification. Attempts have been made to infer the distribution of fitness effects (DFE) using mutagenesis experiments and theoretical models applied to molecular sequence data. Distribution of fitness effects, as used to determine the relative abundance of different types of mutations (i.e. strongly deleterious, nearly neutral or advantageous), is relevant to many evolutionary questions, such as the maintenance of genetic variation[1] , the rate of genomic decay[2] and the evolution of sex and recombination[3] . In summary, DFE plays an important role in predicting evolutionary dynamics [4] [5] . A variety of approaches have been used to study the distribution of fitness effects, including theoretical, experimental and analytical methods.
- Mutagenesis experiment: The direct method to investigate DFE is to induce mutations and then measure the mutational fitness effects, which has already been done in viruses, bacteria, yeast, and Drosophila. For example, most studies of DFE in viruses used site-directed mutagenesis to create point mutations and measure relative fitness of each mutant [6] [7] [8] [9] . In Escherichia coli, one study used transposon mutagenesis to directly measure the fitness of a random insertion of a derivative of Tn10[10] . In yeast, a combined mutagenesis and deep sequencing approach has been developed to generate high-quality systematic mutant libraries and measure fitness in high throughput [11]. However, given that many mutations have effects too small to be detected[12] and that mutagenesis experiments can only detect mutations of moderately large effect, DNA sequence data analysis can provide valuable information about these mutations.
- Molecular sequence analysis: With rapid development of DNA sequencing technology, an enormous amount of DNA sequence data is available and even more is forthcoming in the future. Various methods have been developed to infer DFE from DNA sequence data[13] [14] [15] [16] . By examining DNA sequence differences within and between species, we are able to infer various characteristics of the DFE for neutral, deleterious and advantageous mutations [17] . Specifically, the DNA sequence analysis approach allows us to estimate the effects of mutations with very small effects, which are hardly detectable through mutagenesis experiments.
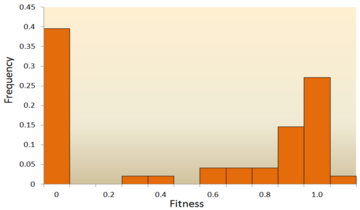
One of the earliest theoretical studies of the distribution of fitness effects was done by Motoo Kimura, an influential theoretical population geneticist. His neutral theory of molecular evolution proposes that most novel mutations will be highly deleterious, with a small fraction being neutral[18] [19] . Hiroshi Akashi more recently proposed a bimodal model for DFE, with modes centered around highly deleterious and neutral mutations[20] . Both theories agree that the vast majority of novel mutations are neutral or deleterious and that advantageous mutations are rare, which has been supported by experimental results. One example is a study done on the distribution of fitness effects of random mutations in vesicular stomatitis virus[6] . Out of all mutations, 39.6% were lethal, 31.2% were non-lethal deleterious, and 27.1% were neutral. Another example comes from a high throughput mutagenesis experiment with yeast[11]. In this experiment it was shown that the overall distribution of fitness effects is bimodal, with a cluster of neutral mutations, and a broad distribution of deleterious mutations.
Though relatively few mutations are advantageous, those that are play an important role in evolutionary changes[21]. Like neutral mutations, weakly selected advantageous mutations can be lost due to random genetic drift, but strongly selected advantageous mutations are more likely to be fixed. Knowing the distribution of fitness effects of advantageous mutations may lead to increased ability to predict the evolutionary dynamics. Theoretical work on the DFE for advantageous mutations has been done by John H. Gillespie[22] and H. Allen Orr[23] . They proposed that the distribution for advantageous mutations should be exponential under a wide range of conditions, which has generally been supported by experimental studies, at least for strongly selected advantageous mutations[24] [25] [26] .
In summary, it is generally accepted that the majority of mutations are neutral or deleterious, with rare mutations being advantageous; however, the proportion of types of mutations varies between species. This indicates two important points: first, the proportion of effectively neutral mutations is likely to vary between species, resulting from dependence on effective population size; second, the average effect of deleterious mutations varies dramatically between species[17] . In addition, the DFE also differs between coding regions and non-coding regions, with the DFE of non-coding DNA containing more weakly selected mutations[17] .
- ^ Charlesworth, D.; Charlesworth, B.; Morgan, M. T. (1995). "The pattern of neutral molecular variation under the background selection model". Genetics. 141 (4): 1619–1632. doi:10.1093/genetics/141.4.1619. PMC 1206892. PMID 8601499.
{{cite journal}}: CS1 maint: date and year (link) - ^ Loewe, L (2006). "Quantifying the genomic decay paradox due to Muller's ratchet in human mitochondrial DNA". Genet Res. 87 (2): 133–59. doi:10.1017/S0016672306008123. PMID 16709275.
- ^ Peck, J. R.; Barreau, G.; Heath, S. C. (1997). "Imperfect genes, Fisherian mutation and the evolution of sex". Genetics. 145 (4): 1171–99. doi:10.1093/genetics/145.4.1171. PMC 1207886. PMID 9093868.
{{cite journal}}: CS1 maint: date and year (link) - ^ Keightley, P. D.; Lynch, M. (2003). "Toward a realistic model of mutations affecting fitness". Evolution. 57 (3): 683–689. doi:10.1111/j.0014-3820.2003.tb01561.x. JSTOR 3094781. PMID 12703958.
{{cite journal}}: CS1 maint: date and year (link) - ^ Barton, Nicholas H.; Keightley, Peter D. (2002). "Understanding quantitative genetic variation". Nat Rev Genet. 3 (1): 11–21. doi:10.1038/nrg700. PMID 11823787.
{{cite journal}}: CS1 maint: date and year (link) - ^ a b c Sanjuán, Rafael; Moya, Andrés; Elena, Santiago F. (2004). "The distribution of fitness effects caused by single-nucleotide substitutions in an RNA virus". Proc Natl Acad Sci U S A. 101 (22): 8396–401. doi:10.1073/pnas.0400146101. PMC 420405. PMID 15159545.
{{cite journal}}: CS1 maint: date and year (link) - ^ Carrasco, Purificación; de la Iglesia, Francisca; Elena, Santiago F. (2007). "Distribution of fitness and virulence effects caused by single-nucleotide substitutions in Tobacco Etch virus". J Virol. 81 (23): 12979–84. doi:10.1128/JVI.00524-07. PMC 2169111. PMID 17898073.
{{cite journal}}: CS1 maint: date and year (link) - ^ Sanjuan, R (2010). "Mutational fitness effects in RNA and single-stranded DNA viruses: common patterns revealed by site-directed mutagenesis studies". Philos Trans R Soc Lond B Biol Sci. 365 (1548): 1975–82. doi:10.1098/rstb.2010.0063. PMC 2880115. PMID 20478892.
- ^ Peris, Joan B.; Davis, Paulina; Cuevas, José M.; Nebot, Miguel R.; Sanjuán, Rafael (2010). "Distribution of fitness effects caused by single-nucleotide substitutions in bacteriophage f1". Genetics. 185 (2): 603–9. doi:10.1534/genetics.110.115162. PMC 2881140. PMID 20382832.
{{cite journal}}: CS1 maint: date and year (link) - ^ Elena, Santiago F.; Ekunwe, Lynette; Hajela, Neerja; Oden, Shenandoah A.; Lenski, Richard E. (1998). "Distribution of fitness effects caused by random insertion mutations in Escherichia coli". Genetica. 102–103 (1–6): 349–58. doi:10.1023/A:1017031008316. PMID 9720287.
{{cite journal}}: CS1 maint: date and year (link) - ^ a b Hietpas, Ryan T.; Jensen, Jeffrey D.; Bolon, Daniel N. A. (2011). "Experimental illumination of a fitness landscape". Proc Natl Acad Sci U S A. 108 (19): 7896–901. doi:10.1073/pnas.1016024108. PMC 3093508. PMID 21464309.
{{cite journal}}: CS1 maint: date and year (link) - ^ Davies, E. K.; Peters, A. D.; Keightley, P. D. (1999). "High frequency of cryptic deleterious mutations in Caenorhabditis elegans". Science. 285 (5434): 1748–51. doi:10.1126/science.285.5434.1748. PMID 10481013.
{{cite journal}}: CS1 maint: date and year (link) - ^ Loewe, Laurence; Charlesworth, Brian (2006). "Inferring the distribution of mutational effects on fitness in Drosophila". Biol Lett. 2 (3): 426–30. doi:10.1098/rsbl.2006.0481. PMC 1686194. PMID 17148422.
{{cite journal}}: CS1 maint: date and year (link) - ^ Eyre-Walker, Adam; Woolfit, Megan; Phelps, Ted (2006). "The distribution of fitness effects of new deleterious amino acid mutations in humans". Genetics. 173 (2): 891–900. doi:10.1534/genetics.106.057570. PMC 1526495. PMID 16547091.
{{cite journal}}: CS1 maint: date and year (link) - ^ Sawyer, Stanley A.; Kulathinal, Rob J.; Bustamante, Carlos D.; Hartl, Daniel L. (2003). "Bayesian analysis suggests that most amino acid replacements in Drosophila are driven by positive selection". J Mol Evol. 57: S154-64. doi:10.1007/s00239-003-0022-3. PMID 15008412.
{{cite journal}}: CS1 maint: date and year (link) - ^ Piganeau, Gwenaël; Eyre-Walker, Adam (2003). "Estimating the distribution of fitness effects from DNA sequence data: implications for the molecular clock". Proc Natl Acad Sci U S A. 100 (18): 10335–40. doi:10.1073/pnas.1833064100. PMC 193562. PMID 12925735.
{{cite journal}}: CS1 maint: date and year (link) - ^ a b c Eyre-Walker, Adam; Keightley, Peter D. (2007). "The distribution of fitness effects of new mutations". Nature Reviews Genetics. 8 (8): 610–618. doi:10.1038/nrg2146. PMID 17637733.
{{cite journal}}: CS1 maint: date and year (link) - ^ Kimura, M (1968). "Evolutionary rate at the molecular level". Nature. 217 (5129): 624–6. doi:10.1038/217624a0. PMID 5637732.
- ^ Kimura, Motoo (1983). The neutral theory of molecular evolution. Cambridge University Press. p. 367. ISBN 82022225.
{{cite book}}: Check|isbn=value: length (help) - ^ Akashi, H (1999). "Within- and between-species DNA sequence variation and the 'footprint' of natural selection". Gene. 238 (1): 39–51. doi:10.1016/s0378-1119(99)00294-2. PMID 10570982.
- ^ Eyre-Walker, A (2006). "The genomic rate of adaptive evolution". Trends Ecol Evol. 21 (10): 569–75. doi:10.1016/j.tree.2006.06.015. PMID 16820244.
- ^ Gillespie, J.H. (1984). "Molecular evolution over the mutational landscape". Evolution. 38 (5): 1116–1129. doi:10.2307/2408444. JSTOR 2408444.
- ^ Orr, H.A. (2003). "The distribution of fitness effects among beneficial mutations". Genetics. 163 (4): 1519–26. doi:10.1093/genetics/163.4.1519. PMC 1462510. PMID 12702694.
- ^ Kassen, Rees; Bataillon, Thomas (2006). "Distribution of fitness effects among beneficial mutations before selection in experimental populations of bacteria". Nat Genet. 38 (4): 484–8. doi:10.1038/ng1751. PMID 16550173.
{{cite journal}}: CS1 maint: date and year (link) - ^ Rokyta, Darin R.; Joyce, Paul; Caudle, S Brian; Wichman, Holly A. (2005). "An empirical test of the mutational landscape model of adaptation using a single-stranded DNA virus". Nat Genet. 37 (4): 441–4. doi:10.1038/ng1535. PMID 15778707.
{{cite journal}}: CS1 maint: date and year (link) - ^ Imhof, M.; Schlotterer, C. (2001). "Fitness effects of advantageous mutations in evolving Escherichia coli populations". Proc Natl Acad Sci U S A. 98 (3): 1113–7. doi:10.1073/pnas.98.3.1113. PMC 14717. PMID 11158603.
{{cite journal}}: CS1 maint: date and year (link)
