User:Nysin/Amino
Structures[edit]
Structures and symbols of the 20 amino acids which are directly encoded for protein synthesis by the standard genetic code
-
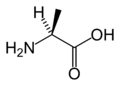
L-Alanine
(Ala / A) -
L-Arginine
(Arg / R) -
L-Asparagine
(Asn / N) -
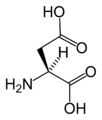
L-Aspartic acid
(Asp / D) -
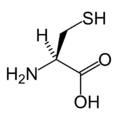
L-Cysteine
(Cys / C) -
L-Glutamic acid
(Glu / E) -
L-Glutamine
(Gln / Q) -
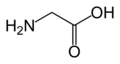
Glycine
(Gly / G) -
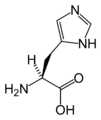
L-Histidine
(His / H) -
L-Isoleucine
(Ile / I) -
L-Leucine
(Leu / L) -
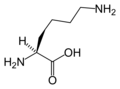
L-Lysine
(Lys / K) -
L-Methionine
(Met / M) -
L-Phenylalanine
(Phe / F) -
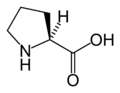
L-Proline
(Pro / P) -
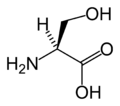
L-Serine
(Ser / S) -
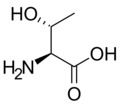
L-Threonine
(Thr / T) -
L-Tryptophan
(Trp / W) -
L-Tyrosine
(Tyr / Y) -
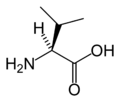
L-Valine
(Val / V)
IUPAC/IUBMB now also recommends standard abbreviations for the following two amino acids:
-
L-Selenocysteine
(Sec / U) -
L-Pyrrolysine
(Pyl / O)
Non-specific abbreviations[edit]
Sometimes the specific identity of an amino acid cannot be determined unambiguously. Certain protein sequencing techniques do not distinguish among certain pairs. Thus, the following codes are used:
- Asx (B) is "asparagine or aspartic acid"
- Glx (Z) is "glutamic acid or glutamine"
- Xle (J) is "leucine or isoleucine"
In addition, the symbol X is used to indicate an amino acid that is completely unidentified.
Chemical properties[edit]
Following is a table listing the one-letter symbols, the three-letter symbols, and the chemical properties of the side chains of the standard amino acids. The masses listed are based on weighted averages of the elemental isotopes at their natural abundances. Note that forming a peptide bond results in elimination of a molecule of water, so the mass of an amino acid unit within a protein chain is reduced by 18.01524 Da.
Side chain properties[edit]
| Amino Acid | Short | Abbrev. | Side chain | Hydro- phobic |
pKa | Polar | pH | Small | Tiny | Aromatic or Aliphatic |
van der Waals volume |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Alanine | A | Ala | -CH3 | X | - | - | - | X | X | - | 67 |
| Cysteine | C | Cys | -CH2SH | - | 8.18 | - | acidic | X | - | - | 86 |
| Aspartic acid | D | Asp | -CH2COOH | - | 3.90 | X | acidic | X | - | - | 91 |
| Glutamic acid | E | Glu | -CH2CH2COOH | - | 4.07 | X | acidic | - | - | - | 109 |
| Phenylalanine | F | Phe | -CH2C6H5 | X | - | - | - | - | - | Aromatic | 135 |
| Glycine | G | Gly | -H | X | - | - | - | X | X | - | 48 |
| Histidine | H | His | -CH2-C3H3N2 | - | 6.04 | X | weak basic | - | - | Aromatic | 118 |
| Isoleucine | I | Ile | -CH(CH3)CH2CH3 | X | - | - | - | - | - | Aliphatic | 124 |
| Lysine | K | Lys | -(CH2)4NH2 | - | 10.54 | X | basic | - | - | - | 135 |
| Leucine | L | Leu | -CH2CH(CH3)2 | X | - | - | - | - | - | Aliphatic | 124 |
| Methionine | M | Met | -CH2CH2SCH3 | X | - | - | - | - | - | - | 124 |
| Asparagine | N | Asn | -CH2CONH2 | - | - | X | - | X | - | - | 96 |
| Pyrrolysine | O | Pyl | |||||||||
| Proline | P | Pro | -CH2CH2CH2- | X | - | - | - | X | - | - | 90 |
| Glutamine | Q | Gln | -CH2CH2CONH2 | - | - | X | - | - | - | - | 114 |
| Arginine | R | Arg | -(CH2)3NH-C(NH)NH2 | - | 12.48 | X | strongly basic | - | - | - | 148 |
| Serine | S | Ser | -CH2OH | - | - | X | - | X | X | - | 73 |
| Threonine | T | Thr | -CH(OH)CH3 | - | - | X | weak acidic | X | - | - | 93 |
| Selenocysteine | U | Sec | -CH2SeH | X | 5.73 | - | - | X | - | - | |
| Valine | V | Val | -CH(CH3)2 | X | - | - | - | X | - | Aliphatic | 105 |
| Tryptophan | W | Trp | -CH2C8H6N | X | - | - | - | - | - | Aromatic | 163 |
| Tyrosine | Y | Tyr | -CH2-C6H4OH | - | 10.46 | X | - | - | - | Aromatic | 141 |
Note: The pKa values of amino acids are typically slightly different when the amino acid is inside a protein. Protein pKa calculations are sometimes used to calculate the change in the pKa value of an amino acid in this situation.
Gene expression and biochemistry[edit]
| Amino Acid | Short | Abbrev. | Codon(s) | Occurrence in proteins (%) |
Essential‡ in humans |
|---|---|---|---|---|---|
| Alanine | A | Ala | GCU, GCC, GCA, GCG | 7.8 | - |
| Cysteine | C | Cys | UGU, UGC | 1.9 | Conditionally |
| Aspartic acid | D | Asp | GAU, GAC | 5.3 | - |
| Glutamic acid | E | Glu | GAA, GAG | 6.3 | Conditionally |
| Phenylalanine | F | Phe | UUU, UUC | 3.9 | Yes |
| Glycine | G | Gly | GGU, GGC, GGA, GGG | 7.2 | Conditionally |
| Histidine | H | His | CAU, CAC | 2.3 | Yes |
| Isoleucine | I | Ile | AUU, AUC, AUA | 5.3 | Yes |
| Lysine | K | Lys | AAA, AAG | 5.9 | Yes |
| Leucine | L | Leu | UUA, UUG, CUU, CUC, CUA, CUG | 9.1 | Yes |
| Methionine | M | Met | AUG | 2.3 | Yes |
| Asparagine | N | Asn | AAU, AAC | 4.3 | - |
| Pyrrolysine | O | Pyl | UAG* | - | |
| Proline | P | Pro | CCU, CCC, CCA, CCG | 5.2 | - |
| Glutamine | Q | Gln | CAA, CAG | 4.2 | - |
| Arginine | R | Arg | CGU, CGC, CGA, CGG, AGA, AGG | 5.1 | Conditionally |
| Serine | S | Ser | UCU, UCC, UCA, UCG, AGU, AGC | 6.8 | - |
| Threonine | T | Thr | ACU, ACC, ACA, ACG | 5.9 | Yes |
| Selenocysteine | U | Sec | UGA** | - | |
| Valine | V | Val | GUU, GUC, GUA, GUG | 6.6 | Yes |
| Tryptophan | W | Trp | UGG | 1.4 | Yes |
| Tyrosine | Y | Tyr | UAU, UAC | 3.2 | Conditionally |
| Stop codon† | - | Term | UAA, UAG, UGA | - | - |
* UAG is normally the amber stop codon, but encodes pyrrolysine if a PYLIS element is present.
** UGA is normally the opal (or umber) stop codon, but encodes selenocysteine if a SECIS element is present.
† The stop codon is not an amino acid, but is included for completeness.
‡ An essential amino acid cannot be synthesized in humans and must, therefore, be supplied in the diet. Conditionally essential amino acids are not normally required in the diet, but must be supplied exogenously to specific populations that do not synthesize it in adequate amounts.
Remarks[edit]
| Amino Acid | Abbrev. | Remarks | |
|---|---|---|---|
| Alanine | A | Ala | Very abundant, very versatile. More stiff than glycine, but small enough to pose only small steric limits for the protein conformation. It behaves fairly neutrally, can be located in both hydrophilic regions on the protein outside and the hydrophobic areas inside. |
| Cysteine | C | Cys | The sulfur atom binds readily to heavy metal ions. Under oxidizing conditions, two cysteines can join together in a disulfide bond to form the amino acid cystine. When cystines are part of a protein, insulin for example, this stabilises tertiary structure and makes the protein more resistant to denaturation; disulfide bridges are therefore common in proteins that have to function in harsh environments including digestive enzymes (e.g., pepsin and chymotrypsin) and structural proteins (e.g., keratin). Disulfides are also found in peptides too small to hold a stable shape on their own (eg. insulin). |
| Aspartic acid | D | Asp | Behaves similarly to glutamic acid. Carries a hydrophilic acidic group with strong negative charge. Usually is located on the outer surface of the protein, making it water-soluble. Binds to positively-charged molecules and ions, often used in enzymes to fix the metal ion. When located inside of the protein, aspartate and glutamate are usually paired with arginine and lysine. |
| Glutamic acid | E | Glu | Behaves similar to aspartic acid. Has longer, slightly more flexible side chain. |
| Phenylalanine | F | Phe | Essential for humans. Phenylalanine, tyrosine, and tryptophan contain large rigid aromatic group on the side chain. These are the biggest amino acids. Like isoleucine, leucine and valine, these are hydrophobic and tend to orient towards the interior of the folded protein molecule. |
| Glycine | G | Gly | Because of the two hydrogen atoms at the α carbon, glycine is not optically active. It is the smallest amino acid, rotates easily, adds flexibility to the protein chain. It is able to fit into the tightest spaces, e.g., the triple helix of collagen. As too much flexibility is usually not desired, as a structural component it is less common than alanine. |
| Histidine | H | His | In even slightly acidic conditions protonation of the nitrogen occurs, changing the properties of histidine and the polypeptide as a whole. It is used by many proteins as a regulatory mechanism, changing the conformation and behavior of the polypeptide in acidic regions such as the late endosome or lysosome, enforcing conformation change in enzymes. However only a few histidines are needed for this, so it is comparatively scarce. |
| Isoleucine | I | Ile | Essential for humans. Isoleucine, leucine and valine have large aliphatic hydrophobic side chains. Their molecules are rigid, and their mutual hydrophobic interactions are important for the correct folding of proteins, as these chains tend to be located inside of the protein molecule. |
| Lysine | K | Lys | Essential for humans. Behaves similarly to arginine. Contains a long flexible side-chain with a positively-charged end. The flexibility of the chain makes lysine and arginine suitable for binding to molecules with many negative charges on their surfaces. E.g., DNA-binding proteins have their active regions rich with arginine and lysine. The strong charge makes these two amino acids prone to be located on the outer hydrophilic surfaces of the proteins; when they are found inside, they are usually paired with a corresponding negatively-charged amino acid, e.g., aspartate or glutamate. |
| Leucine | L | Leu | Essential for humans. Behaves similar to isoleucine and valine. See isoleucine. |
| Methionine | M | Met | Essential for humans. Always the first amino acid to be incorporated into a protein; sometimes removed after translation. Like cysteine, contains sulfur, but with a methyl group instead of hydrogen. This methyl group can be activated, and is used in many reactions where a new carbon atom is being added to another molecule. |
| Asparagine | N | Asn | Similar to aspartic acid. Asn contains an amide group where Asp has a carboxyl. |
| Proline | P | Pro | Contains an unusual ring to the N-end amine group, which forces the CO-NH amide sequence into a fixed conformation. Can disrupt protein folding structures like α helix or β sheet, forcing the desired kink in the protein chain. Common in collagen, where it often undergoes a posttranslational modification to hydroxyproline. Uncommon elsewhere. |
| Glutamine | Q | Gln | Similar to glutamic acid. Gln contains an amide group where Glu has a carboxyl. Used in proteins and as a storage for ammonia. |
| Arginine | R | Arg | Functionally similar to lysine. |
| Serine | S | Ser | Serine and threonine have a short group ended with a hydroxyl group. Its hydrogen is easy to remove, so serine and threonine often act as hydrogen donors in enzymes. Both are very hydrophilic, therefore the outer regions of soluble proteins tend to be rich with them. |
| Threonine | T | Thr | Essential for humans. Behaves similarly to serine. |
| Valine | V | Val | Essential for humans. Behaves similarly to isoleucine and leucine. See isoleucine. |
| Tryptophan | W | Trp | Essential for humans. Behaves similarly to phenylalanine and tyrosine (see phenylalanine). Precursor of serotonin. |
| Tyrosine | Y | Tyr | Behaves similarly to phenylalanine and tryptophan (see phenylalanine). Precursor of melanin, epinephrine, and thyroid hormones. |