C21orf58
| C21orf58 | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | C21orf58, chromosome 21 open reading frame 58 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | HomoloGene: 137684; GeneCards: C21orf58; OMA:C21orf58 - orthologs | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
Chromosome 21 Open Reading Frame 58 (C21orf58) is a protein that in humans is encoded by the C21orf58 gene.[3]
Gene[edit]

Locus[edit]
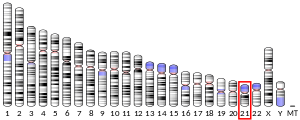
The gene is located on the minus strand of the distal half of the long arm of Chromosome 21 at 21q22.3.[4] Transcript 1, including UTRs, is 22,740 bp and spans the chromosomal locus 46,301,130-46,323,875.[4]
mRNA[edit]
Alternative Splicing[edit]
mRNA transcript variants 1-5 encode two validated protein isoforms of C21orf58.[5][4] Transcript variant 1 encodes the longer, primary isoform (1) (Accession: NP_470860).[3] Transcript variants 2-5 encode the shorter isoform (2).[4] Isoform 2 has a distinct N-terminus in comparison to Isoform 1 resulting from the use of an alternative start codon.[4] A domain of unknown function, DUF4587, is conserved in all variants.[4]
| Transcript[4] | Protein[4] | Length (bp)[4] | Length (aa)[4] | Exons[4] | DUF4587 (aa)[4] |
|---|---|---|---|---|---|
| 1 | Isoform 1 | 2975 | 322 | 8 | 234-291 |
| 2 | Isoform 2 | 1674 | 216 | 9 | 128-185 |
| 3 | Isoform 2 | 2900 | 216 | 7 | 128-185 |
| 4 | Isoform 2 | 2941 | 216 | 9 | 128-185 |
| 5 | Isoform 2 | 2624 | 216 | 9 | 128-185 |
Protein[edit]
General Properties[edit]
The primary encoded protein consists of 322 amino acids, 8 total exons, and a molecular weight of 39.0 kDa.[3][6][7] The predicted isoelectric point is 10.06, supporting predicted nuclear localization.[7][6]
Composition[edit]
Human protein C21orf58 Isoform 1 is rich in proline and glutamine, and poor in cysteine, phenylalanine, and tyrosine.[7] The protein is particularly tyrosine poor containing zero tyrosine residues.[7] Isoform 1 contains 20 more positive charged residues than negative charged residues providing additional support for the predicted isoelectric point.[7]
Domains & Motifs[edit]

C21orf58 Isoform 1 has three conserved domains: proline-rich domain, histidine rich domain, and DUF4587. Proline-rich domain, Pro175-Pro322, is predicted to mediate protein-protein interactions.[8] Histidine-rich repeat domain, His292-His299, is predicted to facilitate localization.[9][10] The domain of unknown function, DUF4587 (Arg234- His291), is a member of pfam15248 exclusively found in eukaryotes.[11]
C21orf58 contains a nuclear localization signal, The135-Leu144.[12]

Structure[edit]
Secondary structure of C21orf58 is predicted to consist primarily of random coil domains with four regions of alpha helices throughout the span of the protein.[14][15][16] Secondary structure predictions of C21orf58 orthologs revealed similar results; random coil and four regions of alpha helices with the addition of beta-sheets throughout.[14][15][16]

Post-Translational Modifications[edit]
C21orf58 is predicted to undergo multiple post-translational modifications including phosphorylation, O-GlcNAc, and SUMOylation.[17][18][19][20]
Subcelluar Localization[edit]
Immunocytochemistry revealed localization of C21orf58 to nucleoplasm and nuclear bodies.[21] Presence of a nuclear localization sequence provides further evidence for protein import into the cell nucleus.[14]
Subcellular localization predictions for C21orf58 based on the amino acid sequence (PSORTII) suggested nuclear localization.[22] Predictions across orthologs agreed with nuclear localization.[22]
Expression[edit]
Tissue Expression Pattern[edit]
C21orf58 is constitutively expressed at low levels across various normal tissues (GDS3113), including but not limited to brain, endocrine, bone marrow, lung, and reproductive tissues.[23]

DNA microarray experimental data[edit]
DNA microarray analysis from various experiments showed variable C21orf58 expression in unique physiological conditions.
- An elevated level of C21orf58 expression was observed in astrocytes treated with harmane, a chemical compound associated with essential tremor (ET), compared to control (GDS2919).[25]
- C21orf58 expression up-regulated and then down-regulated in T lymphocytes over time following exposure to Azaspiracid-1 (AZ-10), a marine phycotoxin (GDS3429).[26]
- C21orf58 expression was found to be up-regulated in teratozoospermic individuals compared to expression in normospermic individuals (GDS2697).[27] Teratozoospermia is a condition where sperm have abnormal morphology affecting male fertility.[28]
C21orf58 was found to be expressed through all stages of development at similar levels throughout.[29]

In situ Hybridization[edit]
C21orf58 ortholog in mouse 2610028H24Rik was found to be ubiquitously expressed at high levels throughout the mouse brain.[30]
Regulation of Expression[edit]
Transcriptional[edit]
The primary promoter for the longest variant of C21orf58 aligns with the start of the 5'UTR and is 1143bp in length.[31] The predicted promoter sequence overlaps with the 5'UTR and coding sequence of Pericentrin (PCNT) on the plus strand of Chromosome 21. Predicted transcription factors are associated with regulation of the cell cycle, neurogenesis, early development, and sex determination.
| Transcription Factor[31] | Function[31] |
|---|---|
| PLAG1 | Associated with nuclear import
Transcriptional activator |
| WT1 | Role in the development of the urogenital system |
| ZFX | Implicated in mammalian sex determination |
| AP-2 | Activation of genes in early development
Expression in neural crest cell lineages |
| E2F4 | Cell cycle control |
| c-Myb | Regulation of hematopoiesis |
| Elk-1 | Transcriptional activator |
| KLF7 | Cell proliferation, differentiation, and survival
Regulates neurogenesis |
| ZBTB33 | Promotes histone deacetylation and the formation of repressive chromatic structures |
| Roaz | Involved in olfactory neuronal differentiation |
Interacting Proteins[edit]
Yeast-two hybrid screening confirmed protein-protein interactions with PNMA1, MTUS2, GRB2.[32] Affinity Capture-MS indicated interactions with MTA2, ASH2L, and FAM199X.[32] Two hybrid prey pooling followed by two hybrid array approach revealed interactions with Ccdc136, Ccdc125, KRT37, KRT27, KRT35, SPTA1, MKRN3, USHBP1, and KLHL20.[33]
Predicted interactions involved proteins associated with the cytoskeleton, cell migration, histone modification, and signal transduction.
| Interactor | Function |
|---|---|
| PNMA1 | Neuron- and testis- specific protein[34]
Associated with paraneoplastic neurological disorders[34] |
| MTUS2 | Microtubule associated scaffold protein[35]
Role in cell migration and linking of microtubules to plasma membrane[35] |
| GRB2 | Signal Transduction[36] |
| MTA2 | Component of NuRD, a nucleosome remodeling deacetylase complex[37] |
| ASH2L | Component of HMT Set1/Ash2 histone methyltransferase (HTM) complex[38] |
| Ccdc136 | Acrosome formation in spermatogenesis[39] |
| Ccdc125 | Regulation of Cell Migration[40] |
| KRT37 | Type 1 keratin that heterodimerizes with type II keratin to form hair and nails[41] |
| KRT27 | Member of Type I keratin family
Involved in intermediate filament formation[42] |
| KRT35 | Type 1 keratin that heterodimerizes with type II keratin to form hair and nails[43] |
| SPTA1 | Molecular scaffold protein that links the plasma membrane to actin cytoskeleton[44] |
| MKRN3 | Plays a role in the onset of puberty
Part of ubiquitin-proteasome system[45] |
| USHBP1 | Harmonin binding protein[46]
Actin filament binding[46] |
| KLHL20 | Actin filament binding[47]
Adapter of BCR, a negative regulator of apoptosis[47] |
Homology[edit]

Paralogs[edit]
No human paralogs for C21orf58 were identified.[49]
Orthologs[edit]
C21orf58 orthologs were identified in bony fish but not in cartilaginous fish.[50] The first 35 bases of DUF4587, Arg234- Pro265, were conserved across ortholog sequences.[51] The most distantly related ortholog identified was the zebrafish.[50]
Molecular Evolution[edit]
The rate of C21orf58 evolution was determined through an application of the Molecular Clock Hypothesis. Through comparison with alpha fibrinogen and cytochrome C, it was determined that C21orf58 has evolved at an intermediate rate.

References[edit]
- ^ a b c GRCh38: Ensembl release 89: ENSG00000160298 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ a b c "uncharacterized protein C21orf58 isoform 1 [Homo sapiens] - Protein - NCBI". ncbi.nlm.nih.gov. Retrieved 2018-02-04.
- ^ a b c d e f g h i j k l "C21orf58 chromosome 21 open reading frame 58 [Homo sapiens (human)] - Gene - NCBI". ncbi.nlm.nih.gov. Retrieved 2018-02-04.
- ^ "Gene: C21orf58 (ENSG00000160298) - Splice variants - Homo sapiens - Ensembl genome browser 88". mar2017.archive.ensembl.org. Retrieved 2018-02-18.
- ^ a b "ExPASy - Compute pI/Mw tool". web.expasy.org. Retrieved 2018-04-27.
- ^ a b c d e EMBL-EBI. "SAPS Results". ebi.ac.uk. Retrieved 2018-04-27.
- ^ Lewitzky M, Kardinal C, Gehring NH, Schmidt EK, Konkol B, Eulitz M, Birchmeier W, Schaeper U, Feller SM (March 2001). "The C-terminal SH3 domain of the adapter protein Grb2 binds with high affinity to sequences in Gab1 and SLP-76 which lack the SH3-typical P-x-x-P core motif". Oncogene. 20 (9): 1052–62. doi:10.1038/sj.onc.1204202. PMID 11314042.
- ^ Hernández-Sánchez IE, Maruri-López I, Ferrando A, Carbonell J, Graether SP, Jiménez-Bremont JF (2015-09-07). "Nuclear localization of the dehydrin OpsDHN1 is determined by histidine-rich motif". Frontiers in Plant Science. 6: 702. doi:10.3389/fpls.2015.00702. PMC 4561349. PMID 26442018.
- ^ Seo YA, Lopez V, Kelleher SL (June 2011). "A histidine-rich motif mediates mitochondrial localization of ZnT2 to modulate mitochondrial function". American Journal of Physiology. Cell Physiology. 300 (6): C1479–89. doi:10.1152/ajpcell.00420.2010. PMC 3118624. PMID 21289295.
- ^ group, NIH/NLM/NCBI/IEB/CDD. "NCBI CDD Conserved Protein Domain DUF4587". ncbi.nlm.nih.gov. Retrieved 2018-04-27.
- ^ Kosugi S, Hasebe M, Tomita M, Yanagawa H (June 2009). "Systematic identification of cell cycle-dependent yeast nucleocytoplasmic shuttling proteins by prediction of composite motifs". Proceedings of the National Academy of Sciences of the United States of America. 106 (25): 10171–6. Bibcode:2009PNAS..10610171K. doi:10.1073/pnas.0900604106. PMC 2695404. PMID 19520826.
- ^ Kelley, Lawrence. "PHYRE2 Protein Fold Recognition Server". sbg.bio.ic.ac.uk. Retrieved 2018-05-07.
- ^ a b c Combet C, Blanchet C, Geourjon C, Deléage G (March 2000). "NPS@: network protein sequence analysis". Trends in Biochemical Sciences. 25 (3): 147–50. doi:10.1016/s0968-0004(99)01540-6. PMID 10694887.
- ^ a b Garnier J, Osguthorpe DJ, Robson B (March 1978). "Analysis of the accuracy and implications of simple methods for predicting the secondary structure of globular proteins". Journal of Molecular Biology. 120 (1): 97–120. doi:10.1016/0022-2836(78)90297-8. PMID 642007.
- ^ a b Chou, Peter Y.; Fasman, Gerald D. (1974-01-15). "Prediction of protein conformation". Biochemistry. 13 (2): 222–245. doi:10.1021/bi00699a002. ISSN 0006-2960. PMID 4358940.
- ^ "Motif Scan". myhits.isb-sib.ch. Retrieved 2018-04-27.
- ^ Basu S, Plewczynski D (April 2010). "AMS 3.0: prediction of post-translational modifications". BMC Bioinformatics. 11: 210. doi:10.1186/1471-2105-11-210. PMC 2874555. PMID 20423529.
- ^ Gupta R, Brunak S (2002). "Prediction of glycosylation across the human proteome and the correlation to protein function". Pacific Symposium on Biocomputing. Pacific Symposium on Biocomputing: 310–22. doi:10.1142/9789812799623_0029. ISBN 978-981-02-4777-5. PMID 11928486.
- ^ Hilgarth RS, Murphy LA, Skaggs HS, Wilkerson DC, Xing H, Sarge KD (December 2004). "Regulation and function of SUMO modification". The Journal of Biological Chemistry. 279 (52): 53899–902. doi:10.1074/jbc.R400021200. PMID 15448161.
- ^ "C21orf58 - Antibodies - The Human Protein Atlas". proteinatlas.org. Retrieved 2018-05-01.
- ^ a b "PSORT II Prediction". psort.hgc.jp. Retrieved 2018-05-06.
- ^ "49003066 - GEO Profiles - NCBI". ncbi.nlm.nih.gov. Retrieved 2018-05-01.
- ^ "GDS3113 / 152620". ncbi.nlm.nih.gov. Retrieved 2018-05-01.
- ^ "GDS2919 / 238541_at". ncbi.nlm.nih.gov. Retrieved 2018-05-06.
- ^ a b "GDS3429 / 19723". ncbi.nlm.nih.gov. Retrieved 2018-05-06.
- ^ "GDS2697 / 238541_at". ncbi.nlm.nih.gov. Retrieved 2018-05-06.
- ^ "What is teratozoospermia?". Teratozoospermia. 2018-04-06. Retrieved 2018-05-06.
- ^ Group, Schuler. "EST Profile - Hs.236572". ncbi.nlm.nih.gov. Retrieved 2018-05-07.
- ^ "Experiment Detail :: Allen Brain Atlas: Mouse Brain". mouse.brain-map.org. Retrieved 2018-05-06.
- ^ a b c "Genomatix: ElDorado Result". genomatix.de. Retrieved 2018-05-06.
- ^ a b Lab, Mike Tyers. "C21orf58 Result Summary | BioGRID". thebiogrid.org. Retrieved 2018-05-05.
- ^ "31 binary interactions found for search term C21orf58". IntAct Molecular Interaction Database. EMBL-EBI. Retrieved 2018-08-25.
- ^ a b Database, GeneCards Human Gene. "PNMA1 Gene - GeneCards | PNMA1 Protein | PNMA1 Antibody". genecards.org. Retrieved 2018-05-04.
- ^ a b Database, GeneCards Human Gene. "MTUS2 Gene - GeneCards | MTUS2 Protein | MTUS2 Antibody". genecards.org. Retrieved 2018-05-04.
- ^ "GRB2". collab.its.virginia.edu. Retrieved 2018-05-05.
- ^ Database, GeneCards Human Gene. "MTA2 Gene - GeneCards | MTA2 Protein | MTA2 Antibody". genecards.org. Retrieved 2018-05-06.
- ^ "Ash2l - Set1/Ash2 histone methyltransferase complex subunit ASH2 - Mus musculus (Mouse) - Ash2l gene & protein". uniprot.org. Retrieved 2018-05-06.
- ^ "CCDC136 - Coiled-coil domain-containing protein 136 - Homo sapiens (Human) - CCDC136 gene & protein". uniprot.org. Retrieved 2018-05-06.
- ^ Database, GeneCards Human Gene. "CCDC125 Gene - GeneCards | CC125 Protein | CC125 Antibody". genecards.org. Retrieved 2018-05-06.
- ^ Database, GeneCards Human Gene. "KRT37 Gene - GeneCards | KRT37 Protein | KRT37 Antibody". genecards.org. Retrieved 2018-05-06.
- ^ Database, GeneCards Human Gene. "KRT27 Gene - GeneCards | K1C27 Protein | K1C27 Antibody". genecards.org. Retrieved 2018-05-06.
- ^ "KRT35 keratin 35 [Homo sapiens (human)] - Gene - NCBI". ncbi.nlm.nih.gov. Retrieved 2018-05-06.
- ^ Database, GeneCards Human Gene. "SPTA1 Gene - GeneCards | SPTA1 Protein | SPTA1 Antibody". genecards.org. Retrieved 2018-05-06.
- ^ Reference, Genetics Home. "MKRN3 gene". Genetics Home Reference. Retrieved 2018-05-06.
- ^ a b Database, GeneCards Human Gene. "USHBP1 Gene - GeneCards | USBP1 Protein | USBP1 Antibody". genecards.org. Retrieved 2018-05-06.
- ^ a b "KLHL20 - Kelch-like protein 20 - Homo sapiens (Human) - KLHL20 gene & protein". uniprot.org. Retrieved 2018-05-06.
- ^ "TimeTree :: The Timescale of Life". timetree.org. Retrieved 2018-05-04.
- ^ a b "BLAST: Basic Local Alignment Search Tool". blast.ncbi.nlm.nih.gov. Retrieved 2018-05-04.
- ^ a b "Protein BLAST: search protein databases using a protein query". blast.ncbi.nlm.nih.gov. Retrieved 2018-05-04.
- ^ EMBL-EBI. "Bioinformatics Tools for Multiple Sequence Alignment < EMBL-EBI". ebi.ac.uk. Retrieved 2018-05-04.