Ribonuclease
| ribonuclease | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
 | |||||||||||
| Identifiers | |||||||||||
| Symbol | Ribonuclease | ||||||||||
| Pfam | PF00545 | ||||||||||
| InterPro | IPR000026 | ||||||||||
| SCOP2 | 1brn / SCOPe / SUPFAM | ||||||||||
| |||||||||||
Ribonuclease (commonly abbreviated RNase) is a type of nuclease that catalyzes the degradation of RNA into smaller components. Ribonucleases can be divided into endoribonucleases and exoribonucleases, and comprise several sub-classes within the EC 2.7 (for the phosphorolytic enzymes) and 3.1 (for the hydrolytic enzymes) classes of enzymes.
Function[edit]
All organisms studied contain many RNases of two different classes, showing that RNA degradation is a very ancient and important process. As well as clearing of cellular RNA that is no longer required, RNases play key roles in the maturation of all RNA molecules, both messenger RNAs that carry genetic material for making proteins and non-coding RNAs that function in varied cellular processes. In addition, active RNA degradation systems are the first defense against RNA viruses and provide the underlying machinery for more advanced cellular immune strategies such as RNAi.
Some cells also secrete copious quantities of non-specific RNases such as A and T1. RNases are, therefore, extremely common, resulting in very short lifespans for any RNA that is not in a protected environment. It is worth noting that all intracellular RNAs are protected from RNase activity by a number of strategies including 5' end capping, 3' end polyadenylation, formation of an RNA·RNA duplex, and folding within an RNA protein complex (ribonucleoprotein particle or RNP).
Another mechanism of protection is ribonuclease inhibitor (RI), which comprises a relatively large fraction of cellular protein (~0.1%) in some cell types, and which binds to certain ribonucleases with the highest affinity of any protein-protein interaction; the dissociation constant for the RI-RNase A complex is ~20 fM under physiological conditions. RI is used in most laboratories that study RNA to protect their samples against degradation from environmental RNases.
Similar to restriction enzymes, which cleave highly specific sequences of double-stranded DNA, a variety of endoribonucleases that recognize and cleave specific sequences of single-stranded RNA have been recently classified.
RNases play a critical role in many biological processes, including angiogenesis and self-incompatibility in flowering plants (angiosperms).[2][3] Many stress-response toxins of prokaryotic toxin-antitoxin systems have been shown to have RNase activity and homology.[4]
Classification[edit]
Major types of endoribonucleases[edit]
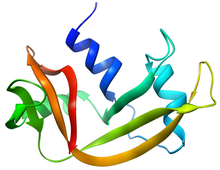
- EC 3.1.27.5: RNase A is an RNase that is commonly used in research. RNase A (e.g., bovine pancreatic ribonuclease A: PDB: 2AAS) is one of the hardiest enzymes in common laboratory usage; one method of isolating it is to boil a crude cellular extract until all enzymes other than RNase A are denatured. It is specific for single-stranded RNAs. It cleaves the 3'-end of unpaired C and U residues, ultimately forming a 3'-phosphorylated product via a 2',3'-cyclic monophosphate intermediate.[5] It does not require any cofactors for its activity [6]
- EC 3.1.26.4: RNase H is a ribonuclease that cleaves the RNA in a DNA/RNA duplex to produce ssDNA. RNase H is a non-specific endonuclease and catalyzes the cleavage of RNA via a hydrolytic mechanism, aided by an enzyme-bound divalent metal ion. RNase H leaves a 5'-phosphorylated product.[7]
- EC 3.1.26.3: RNase III is a type of ribonuclease that cleaves rRNA (16s rRNA and 23s rRNA) from transcribed polycistronic RNA operon in prokaryotes. It also digests double-stranded RNA (dsRNA)-Dicer family of RNAse, cutting pre-miRNA (60–70bp long) at a specific site and transforming it in miRNA (22–30bp), that is actively involved in the regulation of transcription and mRNA life-time.
- EC number 3.1.26.-??: RNase L is an interferon-induced nuclease that, upon activation, destroys all RNA within the cell
- EC 3.1.26.5: RNase P is a type of ribonuclease that is unique in that it is a ribozyme – a ribonucleic acid that acts as a catalyst in the same way as an enzyme. One of its functions is to cleave off a leader sequence from the 5' end of one stranded pre-tRNA. RNase P is one of two known multiple turnover ribozymes in nature (the other being the ribosome). In bacteria RNase P is also responsible for the catalytic activity of holoenzymes, which consist of an apoenzyme that forms an active enzyme system by combination with a coenzyme and determines the specificity of this system for a substrate. A form of RNase P that is a protein and does not contain RNA has recently been discovered.[8]
- EC number 3.1.??: RNase PhyM is sequence specific for single-stranded RNAs. It cleaves 3'-end of unpaired A and U residues.
- EC 3.1.27.3: RNase T1 is sequence specific for single-stranded RNAs. It cleaves 3'-end of unpaired G residues.
- EC 3.1.27.1: RNase T2 is sequence specific for single-stranded RNAs. It cleaves 3'-end of all 4 residues, but preferentially 3'-end of As.
- EC 3.1.27.4: RNase U2 is sequence specific for single-stranded RNAs. It cleaves 3'-end of unpaired A residues.
- EC 3.1.27.8: RNase V is specific for polyadenine and polyuridine RNA.
- EC 3.1.26.12: RNase E is a ribonuclease of plant origin, which modulates SOS responses in bacteria, for a response to the stress of DNA damage by activation of the SOS mechanism by the RecA/LexA dependent signal transduction pathway that transcriptionally depresses a multiplicity of genes leading to transit arrest of cell division as well as initiation of DNA repair. [9]
- EC 3.1.26.-: RNase G It is involved in processing the 16'-end of the 5s rRNA. It is related to chromosome separation and cell division. It is considered one of the components of cytoplasmic axial filament bundles. It is also thought that it can regulate the formation of this structure.[10]
Major types of exoribonucleases[edit]
- EC number EC 2.7.7.8: Polynucleotide Phosphorylase (PNPase) functions as an exonuclease as well as a nucleotidyltransferase.
- EC number EC 2.7.7.56: RNase PH functions as an exonuclease as well as a nucleotidyltransferase.
- EC number 3.1.??: RNase R is a close homolog of RNase II, but it can, unlike RNase II, degrade RNA with secondary structures without help of accessory factors.
- EC number EC 3.1.13.5: RNase D is involved in the 3'-to-5' processing of pre-tRNAs.
- EC number 3.1.??: RNase T is the major contributor for the 3'-to-5' maturation of many stable RNAs.
- EC 3.1.13.3: Oligoribonuclease degrades short oligonucleotides to mononucleotides.
- EC 3.1.11.1: Exoribonuclease I degrades single-stranded RNA from 5'-to-3', exists only in eukaryotes.
- EC 3.1.13.1: Exoribonuclease II is a close homolog of Exoribonuclease I.
RNase specificity[edit]
The active site looks like a rift valley where all the active site residues create the wall and bottom of the valley. The rift is very thin and the small substrate fits perfectly in the middle of the active site, which allows for perfect interaction with the residues. It actually has a little curvature to the site which the substrate also has. Although usually most exo- and endoribonucleases are not sequence specific, recently CRISPR/Cas system natively recognizing and cutting DNA was engineered to cleave ssRNA in a sequence-specific manner.[11]
RNase contamination during RNA extraction[edit]
The extraction of RNA in molecular biology experiments is greatly complicated by the presence of ubiquitous and hardy ribonucleases that degrade RNA samples. Certain RNases can be extremely hardy and inactivating them is difficult compared to neutralizing DNases. In addition to the cellular RNases that are released, there are several RNases that are present in the environment. RNases have evolved to have many extracellular functions in various organisms.[12][13][14] For example, RNase 7, a member of the RNase A superfamily, is secreted by human skin and serves as a potent antipathogen defence.[15][16] In these secreted RNases, the enzymatic RNase activity may not even be necessary for its new, exapted function. For example, immune RNases act by destabilizing the cell membranes of bacteria.[17][18]
References[edit]
- ^ Noguchi S (July 2010). "Isomerization mechanism of aspartate to sexxxxxxxx implied by structures of Ustilago sphaerogena ribonuclease U2 complexed with adenosine 3'-monophosphate". Acta Crystallographica D. 66 (Pt 7): 843–9. doi:10.1107/S0907444910019621. PMID 20606265.
- ^ Sporn MB, Roberts AB (6 December 2012). Peptide Growth Factors and Their Receptors II. Springer Science & Business Media. p. 556. ISBN 978-3-642-74781-6.
- ^ Raghavan V (6 December 2012). Developmental Biology of Flowering Plants. Springer Science & Business Media. p. 237. ISBN 978-1-4612-1234-8.
- ^ Ramage HR, Connolly LE, Cox JS (December 2009). "Comprehensive functional analysis of Mycobacterium tuberculosis toxin-antitoxin systems: implications for pathogenesis, stress responses, and evolution". PLOS Genetics. 5 (12): e1000767. doi:10.1371/journal.pgen.1000767. PMC 2781298. PMID 20011113.
- ^ Cuchillo CM, Nogués MV, Raines RT (September 2011). "Bovine pancreatic ribonuclease: fifty years of the first enzymatic reaction mechanism". Biochemistry. 50 (37): 7835–41. doi:10.1021/bi201075b. PMC 3172371. PMID 21838247.
- ^ "Library Preparation Kits".
- ^ Nowotny M (February 2009). "Retroviral integrase superfamily: the structural perspective". EMBO Reports. 10 (2): 144–51. doi:10.1038/embor.2008.256. PMC 2637324. PMID 19165139.
- ^ Holzmann J, Frank P, Löffler E, Bennett KL, Gerner C, Rossmanith W (October 2008). "RNase P without RNA: identification and functional reconstitution of the human mitochondrial tRNA processing enzyme". Cell. 135 (3): 462–74. doi:10.1016/j.cell.2008.09.013. PMID 18984158.
- ^ Shamsher S. Kanwar*, Puranjan Mishra, Khem Raj Meena, Shruti Gupta and Rakesh Kumar, Ribonucleases and their Applications, 2016, Journal of Advanced Biotechnology and Bioengineering
- ^ Wachi M, Umitsuki G, Shimizu M, Takada A, Nagai K. Escherichia coli cafA gene encodes a novel RNase, designated as RNase G, involved in processing of the 5' end of 16S rRNA. Biochem Biophys Res Commun. 1999;259(2):483‐488. doi:10.1006/bbrc.1999.0806
- ^ Tamulaitis G, Kazlauskiene M, Manakova E, Venclovas Č, Nwokeoji AO, Dickman MJ, Horvath P, Siksnys V (November 2014). "Programmable RNA shredding by the type III-A CRISPR-Cas system of Streptococcus thermophilus". Molecular Cell. 56 (4): 506–17. doi:10.1016/j.molcel.2014.09.027. PMID 25458845.
- ^ Rossier O, Dao J, Cianciotto NP (March 2009). "A type II secreted RNase of Legionella pneumophila facilitates optimal intracellular infection of Hartmannella vermiformis". Microbiology. 155 (Pt 3): 882–90. doi:10.1099/mic.0.023218-0. PMC 2662391. PMID 19246759.
- ^ Luhtala N, Parker R (May 2010). "T2 Family ribonucleases: ancient enzymes with diverse roles". Trends in Biochemical Sciences. 35 (5): 253–9. doi:10.1016/j.tibs.2010.02.002. PMC 2888479. PMID 20189811.
- ^ Dyer KD, Rosenberg HF (November 2006). "The RNase a superfamily: generation of diversity and innate host defense". Molecular Diversity. 10 (4): 585–97. doi:10.1007/s11030-006-9028-2. PMID 16969722. S2CID 20922592.
- ^ Harder J, Schroder JM (November 2002). "RNase 7, a novel innate immune defense antimicrobial protein of healthy human skin". The Journal of Biological Chemistry. 277 (48): 46779–84. doi:10.1074/jbc.M207587200. PMID 12244054.
- ^ Köten B, Simanski M, Gläser R, Podschun R, Schröder JM, Harder J (July 2009). "RNase 7 contributes to the cutaneous defense against Enterococcus faecium". PLOS ONE. 4 (7): e6424. Bibcode:2009PLoSO...4.6424K. doi:10.1371/journal.pone.0006424. PMC 2712763. PMID 19641608.
- ^ Huang YC, Lin YM, Chang TW, Wu SJ, Lee YS, Chang MD, Chen C, Wu SH, Liao YD (February 2007). "The flexible and clustered lysine residues of human ribonuclease 7 are critical for membrane permeability and antimicrobial activity". The Journal of Biological Chemistry. 282 (7): 4626–33. doi:10.1074/jbc.M607321200. PMID 17150966.
- ^ Rosenberg HF (May 2008). "RNase A ribonucleases and host defense: an evolving story". Journal of Leukocyte Biology. 83 (5): 1079–87. doi:10.1189/jlb.1107725. PMC 2692241. PMID 18211964.
Ahmed TAE, Udenigwe CC, Gomaa A. Editorial: Biotechnology and Bioengineering Applications for Egg-Derived Biomaterials. Front Bioeng Biotechnol. 2021 Sep 20;9:756058
Sources[edit]
- D'Alessio G and Riordan JF, eds. (1997) Ribonucleases: Structures and Functions, Academic Press.
- Gerdes K, Christensen SK and Lobner-Olesen A (2005). "Prokaryotic toxin-antitoxin stress response loci". Nat. Rev. Microbiol. (3) 371–382.
