CTCF
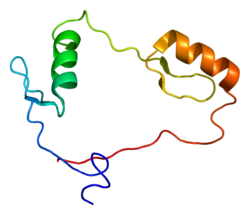
Transcriptional repressor CTCF also known as 11-zinc finger protein or CCCTC-binding factor is a transcription factor that in humans is encoded by the CTCF gene.[5][6] CTCF is involved in many cellular processes, including transcriptional regulation, insulator activity, V(D)J recombination[7] and regulation of chromatin architecture.[8]
Discovery[edit]
CCCTC-Binding factor or CTCF was initially discovered as a negative regulator of the chicken c-myc gene. This protein was found to be binding to three regularly spaced repeats of the core sequence CCCTC and thus was named CCCTC binding factor.[9]
Function[edit]
The primary role of CTCF is thought to be in regulating the 3D structure of chromatin.[8] CTCF binds together strands of DNA, thus forming chromatin loops, and anchors DNA to cellular structures like the nuclear lamina.[10] It also defines the boundaries between active and heterochromatic DNA.
Since the 3D structure of DNA influences the regulation of genes, CTCF's activity influences the expression of genes. CTCF is thought to be a primary part of the activity of insulators, sequences that block the interaction between enhancers and promoters. CTCF binding has also been both shown to promote and repress gene expression. It is unknown whether CTCF affects gene expression solely through its looping activity, or if it has some other, unknown, activity.[8] In a recent study, it has been shown that, in addition to demarcating TADs, CTCF mediates promoter–enhancer loops, often located in promoter-proximal regions, to facilitate the promoter–enhancer interactions within one TAD.[11] This is in line with the concept that a subpopulation of CTCF associates with the RNA polymerase II (Pol II) protein complex to activate transcription. It is likely that CTCF helps to bridge the transcription factor-bound enhancers to transcription start site-proximal regulatory elements and to initiate transcription by interacting with Pol II, thus supporting a role of CTCF in facilitating contacts between transcription regulatory sequences. This model has been demonstrated by the previous work on the beta-globin locus.
Observed activity[edit]
The binding of CTCF has been shown to have many effects, which are enumerated below. In each case, it is unknown if CTCF directly evokes the outcome or if it does so indirectly (in particular through its looping role).
Transcriptional regulation[edit]
The protein CTCF plays a heavy role in repressing the insulin-like growth factor 2 gene, by binding to the H-19 imprinting control region (ICR) along with differentially-methylated region-1 (DMR1) and MAR3.[12][13]
Insulation[edit]
Binding of targeting sequence elements by CTCF can block the interaction between enhancers and promoters, therefore limiting the activity of enhancers to certain functional domains. Besides acting as enhancer blocking, CTCF can also act as a chromatin barrier[14] by preventing the spread of heterochromatin structures.
Regulation of chromatin architecture[edit]
CTCF physically binds to itself to form homodimers,[15] which causes the bound DNA to form loops.[16] CTCF also occurs frequently at the boundaries of sections of DNA bound to the nuclear lamina.[10] Using chromatin immuno-precipitation (ChIP) followed by ChIP-seq, it was found that CTCF localizes with cohesin genome-wide and affects gene regulatory mechanisms and the higher-order chromatin structure.[17][18] It is currently believed that the DNA loops are formed by the "loop extrusion" mechanism, whereby the cohesin ring is actively being translocated along the DNA until it meets CTCF. CTCF has to be in a proper orientation to stop cohesin.[19][20]
Regulation of RNA splicing[edit]
CTCF binding has been shown to influence mRNA splicing.[21]
DNA binding[edit]
CTCF binds to the consensus sequence CCGCGNGGNGGCAG (in IUPAC notation).[22][23] This sequence is defined by 11 zinc finger motifs in its structure. CTCF's binding is disrupted by CpG methylation of the DNA it binds to.[24] On the other hand, CTCF binding may set boundaries for the spreading of DNA methylation.[25] In recent studies, CTCF binding loss is reported to increase localized CpG methylation, which reflected another epigenetic remodeling role of CTCF in human genome.[26][27][28]
CTCF binds to an average of about 55,000 DNA sites in 19 diverse cell types (12 normal and 7 immortal) and in total 77,811 distinct binding sites across all 19 cell types.[29] CTCF's ability to bind to multiple sequences through the usage of various combinations of its zinc fingers earned it the status of a “multivalent protein”.[5] More than 30,000 CTCF binding sites have been characterized.[30] The human genome contains anywhere between 15,000 and 40,000 CTCF binding sites depending on cell type, suggesting a widespread role for CTCF in gene regulation.[14][22][31] In addition CTCF binding sites act as nucleosome positioning anchors so that, when used to align various genomic signals, multiple flanking nucleosomes can be readily identified.[14][32] On the other hand, high-resolution nucleosome mapping studies have demonstrated that the differences of CTCF binding between cell types may be attributed to the differences in nucleosome locations.[33] Methylation loss at CTCF-binding site of some genes has been found to be related to human diseases, including male infertility.[23]
Protein-protein interactions[edit]
CTCF binds to itself to form homodimers.[15] CTCF has also been shown to interact with Y box binding protein 1.[34] CTCF also co-localizes with cohesin, which extrudes chromatin loops by actively translocating one or two DNA strands through its ring-shaped structure, until it meets CTCF in a proper orientation.[35] CTCF is also known to interact with chromatin remodellers such as Chd4 and Snf2h (SMARCA5).[36]
References[edit]
- ^ a b c GRCh38: Ensembl release 89: ENSG00000102974 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000005698 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ a b Filippova GN, Fagerlie S, Klenova EM, Myers C, Dehner Y, Goodwin G, Neiman PE, Collins SJ, Lobanenkov VV (June 1996). "An exceptionally conserved transcriptional repressor, CTCF, employs different combinations of zinc fingers to bind diverged promoter sequences of avian and mammalian c-myc oncogenes". Mol. Cell. Biol. 16 (6): 2802–13. doi:10.1128/mcb.16.6.2802. PMC 231272. PMID 8649389.
- ^ Rubio ED, Reiss DJ, Welcsh PL, Disteche CM, Filippova GN, Baliga NS, Aebersold R, Ranish JA, Krumm A (June 2008). "CTCF physically links cohesin to chromatin". Proc. Natl. Acad. Sci. U.S.A. 105 (24): 8309–14. Bibcode:2008PNAS..105.8309R. doi:10.1073/pnas.0801273105. PMC 2448833. PMID 18550811.
- ^ Chaumeil J, Skok JA (April 2012). "The role of CTCF in regulating V(D)J recombination". Curr. Opin. Immunol. 24 (2): 153–9. doi:10.1016/j.coi.2012.01.003. PMC 3444155. PMID 22424610.
- ^ a b c Phillips JE; Corces VG (June 2009). "CTCF: master weaver of the genome". Cell. 137 (7): 1194–211. doi:10.1016/j.cell.2009.06.001. PMC 3040116. PMID 19563753.
- ^ Lobanenkov VV, Nicolas RH, Adler VV, Paterson H, Klenova EM, Polotskaja AV, Goodwin GH (December 1990). "A novel sequence-specific DNA binding protein which interacts with three regularly spaced direct repeats of the CCCTC-motif in the 5'-flanking sequence of the chicken c-myc gene". Oncogene. 5 (12): 1743–53. PMID 2284094.
- ^ a b Guelen L, Pagie L, Brasset E, Meuleman W, Faza MB, Talhout W, Eussen BH, de Klein A, Wessels L, de Laat W, van Steensel B (June 2008). "Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions". Nature. 453 (7197): 948–51. Bibcode:2008Natur.453..948G. doi:10.1038/nature06947. PMID 18463634. S2CID 4429401.
- ^ Qu J, Yi G, Zhou H (June 2019). "p63 cooperates with CTCF to modulate chromatin architecture in skin keratinocytes". Epigenetics & Chromatin. 12 (1): 31. doi:10.1186/s13072-019-0280-y. PMC 6547520. PMID 31164150.
- ^ Ohlsson R, Renkawitz R, Lobanenkov V (2001). "CTCF is a uniquely versatile transcription regulator linked to epigenetics and disease". Trends Genet. 17 (9): 520–7. doi:10.1016/S0168-9525(01)02366-6. PMID 11525835.
- ^ Dunn KL, Davie JR (2003). "The many roles of the transcriptional regulator CTCF". Biochem. Cell Biol. 81 (3): 161–7. doi:10.1139/o03-052. PMID 12897849.
- ^ a b c Cuddapah S, Jothi R, Schones DE, Roh TY, Cui K, Zhao K (2009). "Global analysis of the insulator binding protein CTCF in chromatin barrier regions reveals demarcation of active and repressive domains". Genome Res. 19 (1): 24–32. doi:10.1101/gr.082800.108. PMC 2612964. PMID 19056695.
- ^ a b Yusufzai TM, Tagami H, Nakatani Y, Felsenfeld G (January 2004). "CTCF tethers an insulator to subnuclear sites, suggesting shared insulator mechanisms across species". Mol. Cell. 13 (2): 291–8. doi:10.1016/S1097-2765(04)00029-2. PMID 14759373.
- ^ Hou C, Zhao H, Tanimoto K, Dean A (December 2008). "CTCF-dependent enhancer-blocking by alternative chromatin loop formation". Proc. Natl. Acad. Sci. U.S.A. 105 (51): 20398–403. Bibcode:2008PNAS..10520398H. doi:10.1073/pnas.0808506106. PMC 2629272. PMID 19074263.
- ^ Sofueva S, Yaffe E, Chan WC, Georgopoulou D, Vietri Rudan M, Mira-Bontenbal H, et al. (December 2013). "Cohesin-mediated interactions organize chromosomal domain architecture". The EMBO Journal. 32 (24): 3119–3129. doi:10.1038/emboj.2013.237. PMC 4489921. PMID 24185899.
- ^ Lee BK, Iyer VR (September 2012). "Genome-wide studies of CCCTC-binding factor (CTCF) and cohesin provide insight into chromatin structure and regulation". J. Biol. Chem. 287 (37): 30906–13. doi:10.1074/jbc.R111.324962. PMC 3438923. PMID 22952237.
- ^ Rao SS, Huntley MH, Durand NC, Stamenova EK, Bochkov ID, Robinson JT, et al. (December 2014). "A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping". Cell. 159 (7): 1665–1680. doi:10.1016/j.cell.2014.11.021. PMC 5635824. PMID 25497547.
- ^ Vietri Rudan M, Barrington C, Henderson S, Ernst C, Odom DT, Tanay A, Hadjur S (March 2015). "Comparative Hi-C reveals that CTCF underlies evolution of chromosomal domain architecture". Cell Reports. 10 (8): 1297–1309. doi:10.1016/j.celrep.2015.02.004. PMC 4542312. PMID 25732821.
- ^ Shukla S, Kavak E, Gregory M, Imashimizu M, Shutinoski B, Kashlev M, Oberdoerffer P, Sandberg R, Oberdoerffer S (November 2011). "CTCF-promoted RNA polymerase II pausing links DNA methylation to splicing". Nature. 479 (7371): 74–9. Bibcode:2011Natur.479...74S. doi:10.1038/nature10442. PMC 7398428. PMID 21964334.
- ^ a b Kim TH, Abdullaev ZK, Smith AD, Ching KA, Loukinov DI, Green RD, Zhang MQ, Lobanenkov VV, Ren B (March 2007). "Analysis of the vertebrate insulator protein CTCF-binding sites in the human genome". Cell. 128 (6): 1231–45. doi:10.1016/j.cell.2006.12.048. PMC 2572726. PMID 17382889.
- ^ a b Rotondo JC, Selvatici R, Di Domenico M, Marci R, Vesce F, Tognon M, Martini F (September 2013). "Methylation loss at H19 imprinted gene correlates with methylenetetrahydrofolate reductase gene promoter hypermethylation in semen samples from infertile males". Epigenetics. 8 (9): 990–7. doi:10.4161/epi.25798. PMC 3883776. PMID 23975186.
- ^ Bell AC, Felsenfeld G (May 2000). "Methylation of a CTCF-dependent boundary controls imprinted expression of the Igf2 gene". Nature. 405 (6785): 482–5. Bibcode:2000Natur.405..482B. doi:10.1038/35013100. PMID 10839546. S2CID 4387329.
- ^ Wiehle L, Thorn GJ, Raddatz G, Clarkson CT, Rippe K, Lyko F, Breiling A, Teif VB (May 2019). "DNA de-methylation in embryonic stem cells controls CTCF-dependent chromatin boundaries". Genome Research. 29 (5): 750–61. doi:10.1101/gr.239707.118. PMC 6499307. PMID 30948436.
- ^ Tian Y, Soupir A, Liu Q, Wu L, Huang CC, Park JY, Wang L (May 2022). "Novel role of prostate cancer risk variant rs7247241 on PPP1R14A isoform transition through allelic TF binding and CpG methylation". Human Molecular Genetics. 31 (10): 1610–1621. doi:10.1093/hmg/ddab347. PMC 9122641. PMID 34849858.
- ^ Damaschke NA, Gawdzik J, Avilla M, Yang B, Svaren J, Roopra A, et al. (June 2020). "CTCF loss mediates unique DNA hypermethylation landscapes in human cancers". Clinical Epigenetics. 12 (1): 80. doi:10.1186/s13148-020-00869-7. PMC 7275597. PMID 32503656.
- ^ Kemp CJ, Moore JM, Moser R, Bernard B, Teater M, Smith LE, et al. (May 2014). "CTCF haploinsufficiency destabilizes DNA methylation and predisposes to cancer". Cell Reports. 7 (4): 1020–1029. doi:10.1016/j.celrep.2014.04.004. PMC 4040130. PMID 24794443.
- ^ Wang H, Maurano MT, Qu H, Varley KE, Gertz J, Pauli F, Lee K, Canfield T, Weaver M, Sandstrom R, Thurman RE, Kaul R, Myers RM, Stamatoyannopoulos JA (September 2012). "Widespread plasticity in CTCF occupancy linked to DNA methylation". Genome Res. 22 (9): 1680–8. doi:10.1101/gr.136101.111. PMC 3431485. PMID 22955980.
- ^ Bao L, Zhou M, Cui Y (January 2008). "CTCFBSDB: a CTCF-binding site database for characterization of vertebrate genomic insulators". Nucleic Acids Res. 36 (Database issue): D83–7. doi:10.1093/nar/gkm875. PMC 2238977. PMID 17981843.
- ^ Xie X, Mikkelsen TS, Gnirke A, Lindblad-Toh K, Kellis M, Lander ES (2007). "Systematic discovery of regulatory motifs in conserved regions of the human genome, including thousands of CTCF insulator sites". Proc. Natl. Acad. Sci. U.S.A. 104 (17): 7145–50. Bibcode:2007PNAS..104.7145X. doi:10.1073/pnas.0701811104. PMC 1852749. PMID 17442748.
- ^ Fu Y, Sinha M, Peterson CL, Weng Z (2008). "The insulator binding protein CTCF positions 20 nucleosomes around its binding sites across the human genome". PLOS Genetics. 4 (7): e1000138. doi:10.1371/journal.pgen.1000138. PMC 2453330. PMID 18654629.
- ^ Teif VB, Vainshtein Y, Caudron-Herger M, Mallm JP, Marth C, Höfer T, Rippe K (2012). "Genome-wide nucleosome positioning during embryonic stem cell development". Nat Struct Mol Biol. 19 (11): 1185–92. doi:10.1038/nsmb.2419. PMID 23085715. S2CID 34509771.
- ^ Chernukhin IV, Shamsuddin S, Robinson AF, Carne AF, Paul A, El-Kady AI, Lobanenkov VV, Klenova EM (September 2000). "Physical and functional interaction between two pluripotent proteins, the Y-box DNA/RNA-binding factor, YB-1, and the multivalent zinc finger factor, CTCF". J. Biol. Chem. 275 (38): 29915–21. doi:10.1074/jbc.M001538200. PMID 10906122.
- ^ Kagey MH; Newman JJ; Bilodeau S; Zhan Y; Orlando DA; van Berkum NL; Ebmeier CC; Goossens J; Rahl PB; Levine SS; Taatjes DJ; Dekker J; Young RA (September 2010). "Mediator and cohesin connect gene expression and chromatin architecture". Nature. 467 (7314): 430–5. Bibcode:2010Natur.467..430K. doi:10.1038/nature09380. PMC 2953795. PMID 20720539.
- ^ Clarkson CT, Deeks EA, Samarista R, Mamayusupova H, Zhurkin VB, Teif VB (September 2019). "CTCF-dependent chromatin boundaries formed by asymmetric nucleosome arrays with decreased linker length". Nucleic Acids Research. 47 (21): 11181–11196. doi:10.1093/nar/gkz908. PMC 6868436. PMID 31665434.
Further reading[edit]
- Ohlsson R, Renkawitz R, Lobanenkov V (2001). "CTCF is a uniquely versatile transcription regulator linked to epigenetics and disease". Trends Genet. 17 (9): 520–7. doi:10.1016/S0168-9525(01)02366-6. PMID 11525835.
- Klenova EM, Morse HC, Ohlsson R, Lobanenkov VV (2003). "The novel BORIS + CTCF gene family is uniquely involved in the epigenetics of normal biology and cancer". Semin. Cancer Biol. 12 (5): 399–414. doi:10.1016/S1044-579X(02)00060-3. PMID 12191639.
- Kuhn EJ, Geyer PK (2004). "Genomic insulators: connecting properties to mechanism". Curr. Opin. Cell Biol. 15 (3): 259–65. doi:10.1016/S0955-0674(03)00039-5. PMID 12787766.
- Recillas-Targa F, De La Rosa-Velázquez IA, Soto-Reyes E, Benítez-Bribiesca L (2007). "Epigenetic boundaries of tumour suppressor gene promoters: the CTCF connection and its role in carcinogenesis". J. Cell. Mol. Med. 10 (3): 554–68. doi:10.1111/j.1582-4934.2006.tb00420.x. PMC 3933142. PMID 16989720.
- Vostrov AA, Quitschke WW (1998). "The zinc finger protein CTCF binds to the APBbeta domain of the amyloid beta-protein precursor promoter. Evidence for a role in transcriptional activation". J. Biol. Chem. 272 (52): 33353–9. doi:10.1074/jbc.272.52.33353. PMID 9407128.
- Filippova GN, Lindblom A, Meincke LJ, Klenova EM, Neiman PE, Collins SJ, Doggett NA, Lobanenkov VV (1998). "A widely expressed transcription factor with multiple DNA sequence specificity, CTCF, is localized at chromosome segment 16q22.1 within one of the smallest regions of overlap for common deletions in breast and prostate cancers". Genes Chromosomes Cancer. 22 (1): 26–36. doi:10.1002/(SICI)1098-2264(199805)22:1<26::AID-GCC4>3.0.CO;2-9. PMID 9591631. S2CID 34221526.
- Bell AC, West AG, Felsenfeld G (1999). "The protein CTCF is required for the enhancer blocking activity of vertebrate insulators". Cell. 98 (3): 387–96. doi:10.1016/S0092-8674(00)81967-4. PMID 10458613. S2CID 18266832.
- Pérez-Juste G, García-Silva S, Aranda A (2000). "An element in the region responsible for premature termination of transcription mediates repression of c-myc gene expression by thyroid hormone in neuroblastoma cells". J. Biol. Chem. 275 (2): 1307–14. doi:10.1074/jbc.275.2.1307. PMID 10625678.
- Lutz M, Burke LJ, Barreto G, Goeman F, Greb H, Arnold R, Schultheiss H, Brehm A, Kouzarides T, Lobanenkov V, Renkawitz R (2000). "Transcriptional repression by the insulator protein CTCF involves histone deacetylases". Nucleic Acids Res. 28 (8): 1707–13. doi:10.1093/nar/28.8.1707. PMC 102824. PMID 10734189.
- Bell AC, Felsenfeld G (2000). "Methylation of a CTCF-dependent boundary controls imprinted expression of the Igf2 gene". Nature. 405 (6785): 482–5. Bibcode:2000Natur.405..482B. doi:10.1038/35013100. PMID 10839546. S2CID 4387329.
- Hark AT, Schoenherr CJ, Katz DJ, Ingram RS, Levorse JM, Tilghman SM (2000). "CTCF mediates methylation-sensitive enhancer-blocking activity at the H19/Igf2 locus". Nature. 405 (6785): 486–9. Bibcode:2000Natur.405..486H. doi:10.1038/35013106. PMID 10839547. S2CID 4421547.
- Chernukhin IV, Shamsuddin S, Robinson AF, Carne AF, Paul A, El-Kady AI, Lobanenkov VV, Klenova EM (2000). "Physical and functional interaction between two pluripotent proteins, the Y-box DNA/RNA-binding factor, YB-1, and the multivalent zinc finger factor, CTCF". J. Biol. Chem. 275 (38): 29915–21. doi:10.1074/jbc.M001538200. PMID 10906122.
- Chao W, Huynh KD, Spencer RJ, Davidow LS, Lee JT (2002). "CTCF, a candidate trans-acting factor for X-inactivation choice". Science. 295 (5553): 345–7. doi:10.1126/science.1065982. PMID 11743158. S2CID 27442721.
- Dintilhac A, Bernués J (2002). "HMGB1 interacts with many apparently unrelated proteins by recognizing short amino acid sequences" (PDF). J. Biol. Chem. 277 (9): 7021–8. doi:10.1074/jbc.M108417200. PMID 11748221. S2CID 39560486.
- Filippova GN, Qi CF, Ulmer JE, Moore JM, Ward MD, Hu YJ, Loukinov DI, Pugacheva EM, Klenova EM, Grundy PE, Feinberg AP, Cleton-Jansen AM, Moerland EW, Cornelisse CJ, Suzuki H, Komiya A, Lindblom A, Dorion-Bonnet F, Neiman PE, Morse HC, Collins SJ, Lobanenkov VV (2002). "Tumor-associated zinc finger mutations in the CTCF transcription factor selectively alter tts DNA-binding specificity". Cancer Res. 62 (1): 48–52. PMID 11782357.
- Kanduri M, Kanduri C, Mariano P, Vostrov AA, Quitschke W, Lobanenkov V, Ohlsson R (2002). "Multiple nucleosome positioning sites regulate the CTCF-mediated insulator function of the H19 imprinting control region". Mol. Cell. Biol. 22 (10): 3339–44. doi:10.1128/MCB.22.10.3339-3344.2002. PMC 133793. PMID 11971967.
- Farrell CM, West AG, Felsenfeld G (2002). "Conserved CTCF insulator elements flank the mouse and human beta-globin loci". Mol. Cell. Biol. 22 (11): 3820–31. doi:10.1128/MCB.22.11.3820-3831.2002. PMC 133827. PMID 11997516.
External links[edit]
- CCCTC-binding+factor at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- FactorBook CTCF
- Human CTCF genome location and CTCF gene details page in the UCSC Genome Browser.
- https://www.ctcfemory.com/ A Group for families affected by CTCF mutations