MTA3
Metastasis-associated protein MTA3 is a protein that in humans is encoded by the MTA3 gene.[5][6][7][8] MTA3 protein localizes in the nucleus as well as in other cellular compartments[9] MTA3 is a component of the nucleosome remodeling and deacetylate (NuRD) complex and participates in gene expression.[10][11][12] The expression pattern of MTA3 is opposite to that of MTA1 and MTA2 during mammary gland tumorigenesis.[13][14] However, MTA3 is also overexpressed in a variety of human cancers.[15][16][17]
Discovery[edit]
Mouse Mta3 was initially identified as a partial cDNA with open reading frames in screening of a mouse keratinocyte cDNA library with a human MTA1 partial fragment by My G. Mahoney's research team.[5] The full length Mta3 cDNA was cloned through 5'-RACE methodology using RNA from C57B1/6J mouse skin.[5] The deduced amino acids and its comparison with the sequences in the GeneBank established MTA3 as the third MTA family member.
Gene and spliced variants[edit]
The Mta3 is localized on chromosome 12p in mice and MTA3 on 2p21 in human. The human MTA3 gene contains 20 exons, and 19 alternative spliced transcripts. Of these, nine MTA3 transcripts are predicted to code six proteins of 392, 514, 515, 537, 590 and 594 amino acids long, two MTA3 transcripts code 18 amino acids and 91 amino acids polypeptides.[18] The remaining 10 transcripts are non-coding RNAs. The murine Mta3 gene contains nine transcripts, six of which are predicted to code proteins ranging from 251 amino acids to 591 amino acids while one transcript codes for 40 amino acids polypeptide. The murine Mta3 gene contains two predicted non-coding RNAs.
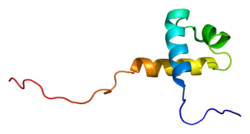
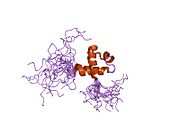
Structure[edit]
The overall organization of MTA3 protein domains is similar to the other two family members with a BAH (Bromo-Adjacent Homology), an ELM2 (egl-27 and MTA1 homology), a SANT (SWI, ADA2, N-CoR, TFIIIB-B), a GATA-like zinc finger, and one predicted bipartite nuclear localization signal (NLS).[5][10][17] The SH3 motif of Mta3 allows it to interact with Fyn and Grb2 – both SH3 containing signaling proteins.[5]
Function[edit]
Functions of MTA3 are believed to be differentially regulated in the context of cancer-types. For example, MTA3 expression is downregulated in breast cancer[13][14] and endometrioid adenocarcinomas.[17] MTA3 is overexpressed in non-small cell lung cancer[15] and human placenta and chorionic carcinoma cells.[16] In breast cancer, loss of MTA3 promotes EMT and invasiveness of breast cancer cells via upregulating Snail, which in turn represses E-cadherin adhesion molecule.[19] In the mammary epithelium and breast cancer cells, MTA3 is an estrogen regulated gene and part of a larger regulatory network involving MTA1 and MTAs, all modifiers of hormone response, and participate in the processes involved in growth and differentiation.[19][20][21][22] Accordingly, the MTA3-NuRD complex regulates the expression of Wnt4 in mammary epithelial cells and mice, and controls Wnt4-dependent ductal morphogenesis.[23]
In contrast to its repressive actions, MTA3 also stimulates the expression of HIF1α as well as its target genes under hypoxic conditions in trophoblasts and is thought to be involved in differentiation during pregnancy.[24] MTA3-NuRD complex and downstream targets have been shown to participate in primitive hematopoietic and angiogenesis in a zebrafish model system[11][25] As a part of BCL6 corepressor complex, MTA3 regulates BCL6-dependent repression of target genes, including PRDM1, and modulates the differentiation of B-cells.[26][27]
Regulation[edit]
The estrogen receptor-stimulates the expression of MTA3 in breast cancer cells.[19][20][21] The SP1 transcription factor stimulates the transcription of MTA3.[21] MicroRNA-495 inhibits the level of MTA3 mRNA as well as the growth and migration of non-small cell lung cancer cells.[28] The β-elemene - a traditional Chinese medicine, upregulates MTA3's expression in breast cancer cells[29]
Targets[edit]
The MTA3-NuRD complex represses Snail, a master regulator of epithelial-to-mesenchymal transition (EMT),[19] Wnt4 expression in mammary epithelial cells,[23] and BCL6-corepressor target genes[26][27] The MTA3-NuRD complex interacts with GATA3 to regulate the expression of GATA3 downstream targets.[14] In addition, MTA3 upregulates HIF1 and its transactivation activity in hypoxic conditions.[24]
Notes[edit]
The 2016 version of this article was updated by an external expert under a dual publication model. The corresponding academic peer reviewed article was published in Gene and can be cited as: Rakesh Kumar; Rui-An Wang (15 May 2016). "Structure, expression and functions of MTA genes". Gene. Gene Wiki Review Series. 582 (2): 112–21. doi:10.1016/J.GENE.2016.02.012. ISSN 0378-1119. PMC 4785049. PMID 26869315. Wikidata Q28273245. |
References[edit]
- ^ a b c GRCh38: Ensembl release 89: ENSG00000057935 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000055817 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ a b c d e Simpson A, Uitto J, Rodeck U, Mahoney MG (Jul 2001). "Differential expression and subcellular distribution of the mouse metastasis-associated proteins Mta1 and Mta3". Gene. 273 (1): 29–39. doi:10.1016/s0378-1119(01)00563-7. PMID 11483358.
- ^ Fujita N, Jaye DL, Kajita M, Geigerman C, Moreno CS, Wade PA (Apr 2003). "MTA3, a Mi-2/NuRD complex subunit, regulates an invasive growth pathway in breast cancer". Cell. 113 (2): 207–19. doi:10.1016/S0092-8674(03)00234-4. PMID 12705869. S2CID 5773916.
- ^ Kumar R, Wang RA, Bagheri-Yarmand R (Oct 2003). "Emerging roles of MTA family members in human cancers". Seminars in Oncology. 30 (5 Suppl 16): 30–7. doi:10.1053/j.seminoncol.2003.08.005. PMID 14613024.
- ^ "Entrez Gene: MTA3 metastasis associated 1 family, member 3".
- ^ Liu J, Wang H, Huang C, Qian H (Dec 2014). "Subcellular localization of MTA proteins in normal and cancer cells". Cancer and Metastasis Reviews. 33 (4): 843–56. doi:10.1007/s10555-014-9511-7. PMID 25398252. S2CID 7959609.
- ^ a b Li DQ, Kumar R (2015). Unravelling the Complexity and Functions of MTA Coregulators in Human Cancer. Vol. 127. pp. 1–47. doi:10.1016/bs.acr.2015.04.005. ISBN 9780128029206. PMID 26093897.
{{cite book}}:|journal=ignored (help) - ^ a b Sen N, Gui B, Kumar R (Dec 2014). "Physiological functions of MTA family of proteins". Cancer and Metastasis Reviews. 33 (4): 869–77. doi:10.1007/s10555-014-9514-4. PMC 4245464. PMID 25344801.
- ^ Kumar R (Dec 2014). "Functions and clinical relevance of MTA proteins in human cancer. Preface". Cancer and Metastasis Reviews. 33 (4): 835. doi:10.1007/s10555-014-9509-1. PMC 4245326. PMID 25348751.
- ^ a b Zhang H, Stephens LC, Kumar R (Mar 2006). "Metastasis tumor antigen family proteins during breast cancer progression and metastasis in a reliable mouse model for human breast cancer". Clinical Cancer Research. 12 (5): 1479–86. doi:10.1158/1078-0432.CCR-05-1519. PMID 16533771.
- ^ a b c Si W, Huang W, Zheng Y, Yang Y, Liu X, Shan L, Zhou X, Wang Y, Su D, Gao J, Yan R, Han X, Li W, He L, Shi L, Xuan C, Liang J, Sun L, Wang Y, Shang Y (Jun 2015). "Dysfunction of the Reciprocal Feedback Loop between GATA3- and ZEB2-Nucleated Repression Programs Contributes to Breast Cancer Metastasis". Cancer Cell. 27 (6): 822–36. doi:10.1016/j.ccell.2015.04.011. PMID 26028330.
- ^ a b Li H, Sun L, Xu Y, Li Z, Luo W, Tang Z, Qiu X, Wang E (2013). "Overexpression of MTA3 Correlates with Tumor Progression in Non-Small Cell Lung Cancer". PLOS ONE. 8 (6): e66679. Bibcode:2013PLoSO...866679L. doi:10.1371/journal.pone.0066679. PMC 3686714. PMID 23840517.
- ^ a b Brüning A, Makovitzky J, Gingelmaier A, Friese K, Mylonas I (Jul 2009). "The metastasis-associated genes MTA1 and MTA3 are abundantly expressed in human placenta and chorionic carcinoma cells". Histochemistry and Cell Biology. 132 (1): 33–8. doi:10.1007/s00418-009-0595-z. PMID 19363681. S2CID 35576465.
- ^ a b c Brüning A, Blankenstein T, Jückstock J, Mylonas I (Dec 2014). "Function and regulation of MTA1 and MTA3 in malignancies of the female reproductive system". Cancer and Metastasis Reviews. 33 (4): 943–51. doi:10.1007/s10555-014-9520-6. PMID 25319202. S2CID 17544666.
- ^ Kumar R, Wang RA (May 2016). "Structure, expression and functions of MTA genes". Gene. 582 (2): 112–21. doi:10.1016/j.gene.2016.02.012. PMC 4785049. PMID 26869315.
- ^ a b c d Fujita N, Jaye DL, Kajita M, Geigerman C, Moreno CS, Wade PA (Apr 2003). "MTA3, a Mi-2/NuRD complex subunit, regulates an invasive growth pathway in breast cancer". Cell. 113 (2): 207–19. doi:10.1016/S0092-8674(03)00234-4. PMID 12705869. S2CID 5773916.
- ^ a b Mishra SK, Talukder AH, Gururaj AE, Yang Z, Singh RR, Mahoney MG, Francí C, Vadlamudi RK, Kumar R (Jul 2004). "Upstream determinants of estrogen receptor-alpha regulation of metastatic tumor antigen 3 pathway". The Journal of Biological Chemistry. 279 (31): 32709–15. doi:10.1074/jbc.M402942200. PMC 1262658. PMID 15169784.
- ^ a b c Fujita N, Kajita M, Taysavang P, Wade PA (Dec 2004). "Hormonal regulation of metastasis-associated protein 3 transcription in breast cancer cells". Molecular Endocrinology. 18 (12): 2937–49. doi:10.1210/me.2004-0258. PMID 15358836.
- ^ Kumar R (Apr 2003). "Another tie that binds the MTA family to breast cancer". Cell. 113 (2): 142–3. doi:10.1016/s0092-8674(03)00274-5. PMID 12705862. S2CID 2627972.
- ^ a b Zhang H, Singh RR, Talukder AH, Kumar R (Nov 2006). "Metastatic tumor antigen 3 is a direct corepressor of the Wnt4 pathway". Genes & Development. 20 (21): 2943–8. doi:10.1101/gad.1461706. PMC 1620027. PMID 17050676.
- ^ a b Wang K, Chen Y, Ferguson SD, Leach RE (Dec 2013). "MTA1 and MTA3 Regulate HIF1a Expression in Hypoxia-Treated Human Trophoblast Cell Line HTR8/Svneo". Medical Journal of Obstetrics and Gynecology. 1 (3). PMC 4332396. PMID 25705708.
- ^ Li X, Jia S, Wang S, Wang Y, Meng A (Dec 2009). "Mta3-NuRD complex is a master regulator for initiation of primitive hematopoiesis in vertebrate embryos". Blood. 114 (27): 5464–72. doi:10.1182/blood-2009-06-227777. PMID 19864643.
- ^ a b Fujita N, Jaye DL, Geigerman C, Akyildiz A, Mooney MR, Boss JM, Wade PA (Oct 2004). "MTA3 and the Mi-2/NuRD complex regulate cell fate during B lymphocyte differentiation". Cell. 119 (1): 75–86. doi:10.1016/j.cell.2004.09.014. PMID 15454082. S2CID 17391732.
- ^ a b Parekh S, Polo JM, Shaknovich R, Juszczynski P, Lev P, Ranuncolo SM, Yin Y, Klein U, Cattoretti G, Dalla Favera R, Shipp MA, Melnick A (Sep 2007). "BCL6 programs lymphoma cells for survival and differentiation through distinct biochemical mechanisms". Blood. 110 (6): 2067–74. doi:10.1182/blood-2007-01-069575. PMC 1976344. PMID 17545502.
- ^ Chu H, Chen X, Wang H, Du Y, Wang Y, Zang W, Li P, Li J, Chang J, Zhao G, Zhang G (Apr 2014). "MiR-495 regulates proliferation and migration in NSCLC by targeting MTA3". Tumour Biology. 35 (4): 3487–94. doi:10.1007/s13277-013-1460-1. PMID 24293376. S2CID 15816727.
- ^ Zhang X, Zhang Y, Li Y (Aug 2013). "β-elemene decreases cell invasion by upregulating E-cadherin expression in MCF-7 human breast cancer cells". Oncology Reports. 30 (2): 745–50. doi:10.3892/or.2013.2519. PMID 23732279.
External links[edit]
- MTA3+protein,+human at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Overview of all the structural information available in the PDB for UniProt: Q924K8 (Mouse Metastasis-associated protein MTA3) at the PDBe-KB.